
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,431,465 – 25,431,576 |
| Length | 111 |
| Max. P | 0.908590 |

| Location | 25,431,465 – 25,431,562 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 76.37 |
| Mean single sequence MFE | -27.84 |
| Consensus MFE | -13.20 |
| Energy contribution | -13.53 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.47 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.898910 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
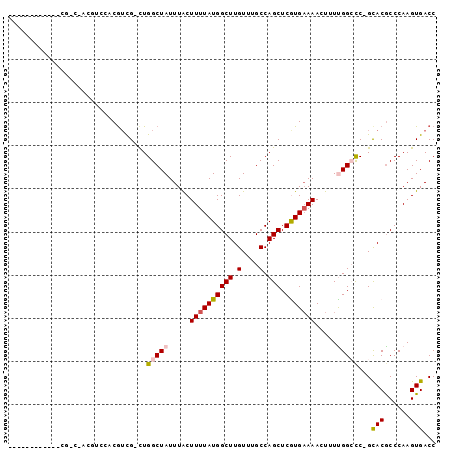
>3R_DroMel_CAF1 25431465 97 - 27905053 AAAAUAUGAGCUCAGCCAGCUCCACGUUG-CUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGGGCC-GCACGCCCAAGUGACC ....(((((((..(((((((........)-))))))...........((((......))))))))))).........((((.-....))))........ ( -35.70) >DroVir_CAF1 137395 79 - 1 ------------CGU------CAUCGUCG-CUGGCUAUUUACUUUUAUGGCUUGUUCGCCAGCUCAUGAAAACUUUUGGCCC-GCACGCCCAAGUGACC ------------.((------((((((.(-(.(((((.....((((((((((.(....).))).))))))).....))))).-))))).....))))). ( -25.00) >DroEre_CAF1 121553 95 - 1 AAGAUACGAGCUCA-C-UCUACCACGUUG-CUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGGGCC-GCACGCCCAAGUGACC ...........(((-(-(....((((..(-(((((......((.....)).......)))))).)))).........((((.-....)))).))))).. ( -27.42) >DroWil_CAF1 146499 80 - 1 ------------------CGUCAUCGACG-UUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGCAAACUUUUGGCCCUGCACACCCAAGUGACC ------------------.(((((.((.(-(((((......((.....)).......))))))))(((((............)))))......))))). ( -19.42) >DroMoj_CAF1 136119 84 - 1 ------------CGUCUAUGCCGUCGUCG-CUGGCUAUUUGCUUUUAUGGCUUGUUUGCCAGCUCAUGAAAACU-UUGGCUC-GUACGCCCAAGUGACC ------------.(((.......((((.(-(((((.((..(((.....)))..))..))))))..))))..(((-(.(((..-....))).))))))). ( -26.70) >DroAna_CAF1 123722 92 - 1 ------CGUGCUUGGCCAGGUUCACAUUGCCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAACUUUUGGCCC-GCACGCCCAAGUGACC ------.(((..(((((((((.......)))))))))..)))......(((.(((..(((((...((....))..)))))..-))).)))......... ( -32.80) >consensus ____________CG_C_ACGUCCACGUCG_CUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAACUUUUGGCCC_GCACGCCCAAGUGACC ................................(((((.....((((((((((.(....).))).))))))).....)))))...(((......)))... (-13.20 = -13.53 + 0.33)


| Location | 25,431,484 – 25,431,576 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 93.53 |
| Mean single sequence MFE | -22.81 |
| Consensus MFE | -17.90 |
| Energy contribution | -17.90 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.908590 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25431484 92 + 27905053 CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGAGCUGGCUGAGCUCAUAUUUUCCA--CAGCAACUGAA ..............((((((((......)))...........((((((((........)))))))))))))..........--(((...))).. ( -26.50) >DroSec_CAF1 114337 94 + 1 CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGAGCUAGCUGAGCUCAUAUUUUCCACACAGCAACUGAA ..((((.....((((.((((((......................))))))..))))(((((.....)))))....))))....(((...))).. ( -21.75) >DroSim_CAF1 116336 92 + 1 CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGAGCUAGCUGAGCUCAUAUUUUCCA--CAGCAACUGAA ..((((.....((((.((((((......................))))))..))))(((((.....)))))....))))..--(((...))).. ( -21.75) >DroEre_CAF1 121572 90 + 1 CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGUAGA-G-UGAGCUCGUAUCUUGCA--CGGCAACUGAA (((..((((((((((.((((((......................))))))..))))).)))-)-)..((.........)).--)))........ ( -22.35) >DroYak_CAF1 124515 91 + 1 CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGAAGU-GCUGAGCUCAUAUUUUCCA--CAGCAACUCAA ..((...((((((((.((((((......................))))))..))))))))(-((((...............--)))))..)).. ( -21.71) >consensus CCGAAACUUUUCACGAGCUGGCAAACAAGCCAUAAAAGUAAAUAGCCAGCAACGUGGAGCU_GCUGAGCUCAUAUUUUCCA__CAGCAACUGAA ........(((((((.((((((......................))))))..)))))))...((((.................))))....... (-17.90 = -17.90 + 0.00)



| Location | 25,431,484 – 25,431,576 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 93.53 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -21.12 |
| Energy contribution | -21.84 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25431484 92 - 27905053 UUCAGUUGCUG--UGGAAAAUAUGAGCUCAGCCAGCUCCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ...........--..((((.(((((((..(((((((........)))))))...........((((......))))))))))).....)))).. ( -29.40) >DroSec_CAF1 114337 94 - 1 UUCAGUUGCUGUGUGGAAAAUAUGAGCUCAGCUAGCUCCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ...............((((.(((((((..(((((((........)))))))...........((((......))))))))))).....)))).. ( -27.10) >DroSim_CAF1 116336 92 - 1 UUCAGUUGCUG--UGGAAAAUAUGAGCUCAGCUAGCUCCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ...........--..((((.(((((((..(((((((........)))))))...........((((......))))))))))).....)))).. ( -27.10) >DroEre_CAF1 121572 90 - 1 UUCAGUUGCCG--UGCAAGAUACGAGCUCA-C-UCUACCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ........(((--.((.......(((....-)-))...((((..((((((......((.....)).......)))))).))))....))..))) ( -21.12) >DroYak_CAF1 124515 91 - 1 UUGAGUUGCUG--UGGAAAAUAUGAGCUCAGC-ACUUCCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ..((((((..(--(((((....((....))..-..))))))((.((.((((((........)))))).))..)))))))).............. ( -23.30) >consensus UUCAGUUGCUG__UGGAAAAUAUGAGCUCAGC_AGCUCCACGUUGCUGGCUAUUUACUUUUAUGGCUUGUUUGCCAGCUCGUGAAAAGUUUCGG ....................(((((((..(((((((........)))))))...........((((......)))))))))))........... (-21.12 = -21.84 + 0.72)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:38:36 2006