
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,138,020 – 3,138,131 |
| Length | 111 |
| Max. P | 0.670181 |

| Location | 3,138,020 – 3,138,131 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 72.62 |
| Mean single sequence MFE | -29.47 |
| Consensus MFE | -17.83 |
| Energy contribution | -17.73 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.670181 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
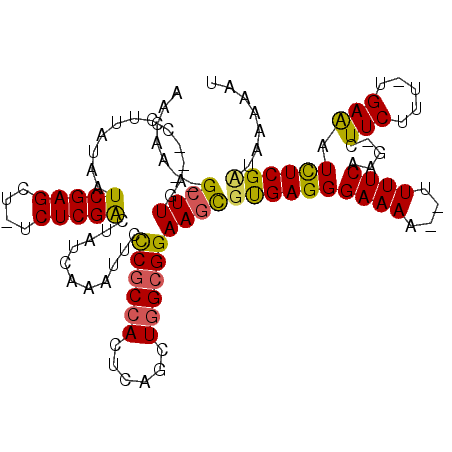
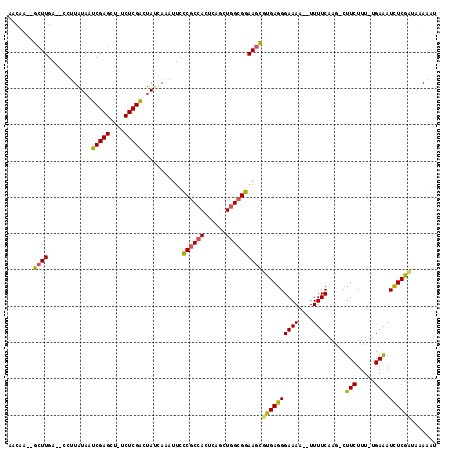
>3R_DroMel_CAF1 3138020 111 - 27905053 AACAA--GCUUGA--CGUUAUAAUCGAGCU-UCUCGGCUAUCAAAUUCCCGCCACUCAGCUGGCGGAAGCGCGAGGGAAAAACUUUUCAAA-CUUCUUU-UGAAAUCUCGUAAGAAAU ...((--((((((--........)))))))-)(((((((.........((((((......)))))).))).)))).........(((((((-.....))-)))))..((....))... ( -28.80) >DroSec_CAF1 13891 117 - 1 AGAGAGAGAUUGUCACUUUGUAAUCGAGCUUUCUCGACUAUUAGAUUCCCGCGACUCAUCUGGAGGAA-CGUGAGGGAAAACGUUUUCAAGUCUUCUUUGUGAGAUUUCGCUAUAAAA .(((((((.(((............))).)))))))...............(((((((((..((((((.-..((((((......))))))..))))))..)))))...))))....... ( -30.70) >DroSim_CAF1 14635 109 - 1 AACAA--GCUUGA--ACGUUUAAUCGAGCU-GCUCGACAAUGAAAUUAUCGCCACUCAGCUGGCGGAAGUAUGAGGGAAAC--UUUUCUAG-CUUCUUU-CGAAAUCUCGAUAAAGGU .....--......--((.((((.(((((.(-..((((............(((((......)))))(((((..((((....)--)))....)-))))..)-))).).))))))))).)) ( -28.90) >consensus AACAA__GCUUGA__CCUUAUAAUCGAGCU_UCUCGACUAUCAAAUUCCCGCCACUCAGCUGGCGGAAGCGUGAGGGAAAA__UUUUCAAG_CUUCUUU_UGAAAUCUCGAUAAAAAU .......((((............(((((....)))))...........((((((......))))))))))((((((((((....)))).....(((.....))).))))))....... (-17.83 = -17.73 + -0.10)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:56:36 2006