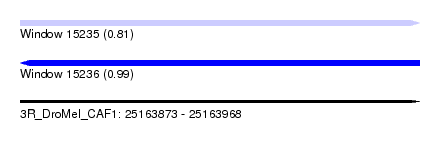

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,163,873 – 25,163,968 |
| Length | 95 |
| Max. P | 0.991805 |

| Location | 25,163,873 – 25,163,968 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 77.59 |
| Mean single sequence MFE | -33.18 |
| Consensus MFE | -15.04 |
| Energy contribution | -16.57 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
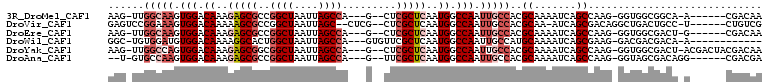
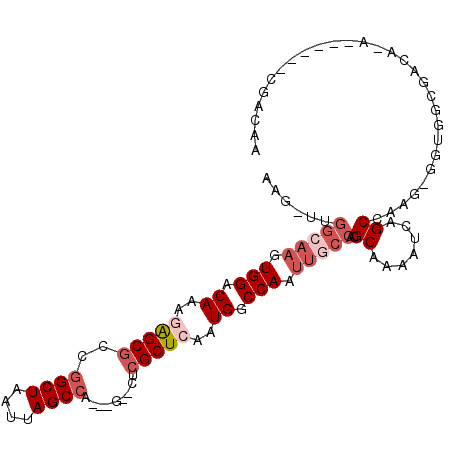
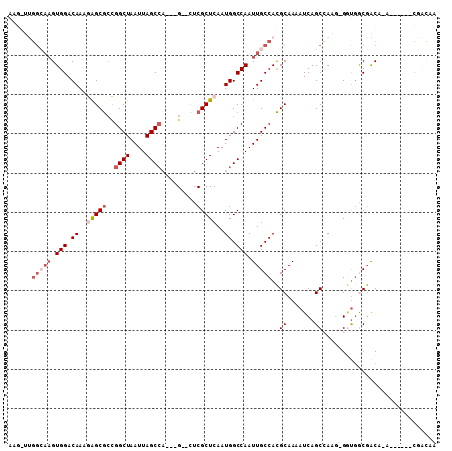
>3R_DroMel_CAF1 25163873 95 + 27905053 AAG-UUGGCAAGUGGACAAAGAGCGCCGGCUAAUUAGCCA---G--CUCGCUCAAUGGCCAAUUGCCACGCAAAAUCAGCCAAG-GGUGGCGGCA-A------CGACAA ..(-(((((((.(((.((..((((((.((((....)))).---)--)..))))..)).))).)))))).))....((.((((..-..))))(...-.------)))... ( -35.00) >DroVir_CAF1 26015 97 + 1 GAGUCCGGAAAGUGGACAAAAAGCGCCGGCUAAUUAGC--CUCG--CUCGCUCAAUGGCCAAUUGCCACGCAA-AUCAGCGACAGGCUGACUGCC-U------CUGUCG ..(((((.....))))).....(((..(((...(((((--((.(--.(((((...((((.....)))).....-...)))))))))))))..)))-.------.))).. ( -30.00) >DroEre_CAF1 23467 95 + 1 AAG-UUGGCAAGUGGACAAAGAGCGCCGGCUAAUUAGCCA---G--CUCGCUCAAUGGCCAAUUGCCACGCAAAAUCAGCCAAG-GGUGGCGACU-G------CGACAA ...-.((((((.(((.((..((((((.((((....)))).---)--)..))))..)).))).))))))((((...((.((((..-..)))))).)-)------)).... ( -37.10) >DroWil_CAF1 98400 91 + 1 GGC-UGUGGAUGUGGACAAAAGGCACUGGCUAAUUAGCCA---GUGUUCGCUCAAUGGCCAAUUGCCAUGCAAAAUCAGCGAAG-GACGACGACA-A------------ ...-(((.(...((..(.....(((((((((....)))))---))))(((((..(((((.....)))))........))))).)-..)).).)))-.------------ ( -29.50) >DroYak_CAF1 23012 101 + 1 AAG-UUGGCCAGUGGACAAAGAGCGGCGGCUAAUUAGCCA---G--CUCGCUCAAUGGCCAAUUGCCACGCAAAAUCAGCCAAG-GGUGGCGACU-ACGACUACGACAA .((-((((((((....)...(((((((((((....)))).---)--).)))))..)))))))))(((((.(............)-.)))))....-............. ( -38.40) >DroAna_CAF1 20725 94 + 1 --U-GUGCCAAGUGGACAAAGAGCGCCGGCUAAUUAGCCA---G--UUCGCUCAAUGGCCAAUUGCCACGCAAAAUCAGCCAAG-GGUAGCGACAGG------CGACGA --(-((.((....)))))....(((((((((....)))).---(--(.(((((..((((...((((...)))).....))))..-.).)))))).))------)).).. ( -29.10) >consensus AAG_UUGGCAAGUGGACAAAGAGCGCCGGCUAAUUAGCCA___G__CUCGCUCAAUGGCCAAUUGCCACGCAAAAUCAGCCAAG_GGUGGCGACA_A______CGACAA ......(((((.(((.((..(((((..((((....)))).........)))))..)).))).)))))..((.......))............................. (-15.04 = -16.57 + 1.53)
| Location | 25,163,873 – 25,163,968 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 77.59 |
| Mean single sequence MFE | -37.50 |
| Consensus MFE | -22.99 |
| Energy contribution | -23.97 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.28 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.61 |
| SVM decision value | 2.29 |
| SVM RNA-class probability | 0.991805 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
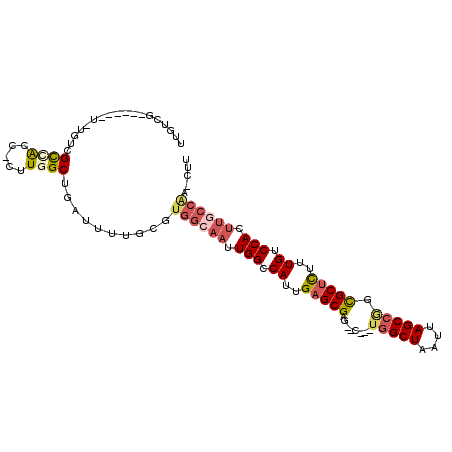
>3R_DroMel_CAF1 25163873 95 - 27905053 UUGUCG------U-UGCCGCCACC-CUUGGCUGAUUUUGCGUGGCAAUUGGCCAUUGAGCGAG--C---UGGCUAAUUAGCCGGCGCUCUUUGUCCACUUGCCAA-CUU ....((------(-..(.((((..-..)))).).....)))((((((.(((.((..((((..(--(---(((((....)))))))))))..)).))).)))))).-... ( -39.30) >DroVir_CAF1 26015 97 - 1 CGACAG------A-GGCAGUCAGCCUGUCGCUGAU-UUGCGUGGCAAUUGGCCAUUGAGCGAG--CGAG--GCUAAUUAGCCGGCGCUUUUUGUCCACUUUCCGGACUC ....((------(-((((((((((.....))))))-).((.((((((((((((.(((......--))))--))))))).)))))))))))).((((.......)))).. ( -36.60) >DroEre_CAF1 23467 95 - 1 UUGUCG------C-AGUCGCCACC-CUUGGCUGAUUUUGCGUGGCAAUUGGCCAUUGAGCGAG--C---UGGCUAAUUAGCCGGCGCUCUUUGUCCACUUGCCAA-CUU ....((------(-((((((((..-..)))).)))..))))((((((.(((.((..((((..(--(---(((((....)))))))))))..)).))).)))))).-... ( -43.90) >DroWil_CAF1 98400 91 - 1 ------------U-UGUCGUCGUC-CUUCGCUGAUUUUGCAUGGCAAUUGGCCAUUGAGCGAACAC---UGGCUAAUUAGCCAGUGCCUUUUGUCCACAUCCACA-GCC ------------.-..........-....((((..((((((((((.....)))))...)))))(((---(((((....)))))))).................))-)). ( -28.00) >DroYak_CAF1 23012 101 - 1 UUGUCGUAGUCGU-AGUCGCCACC-CUUGGCUGAUUUUGCGUGGCAAUUGGCCAUUGAGCGAG--C---UGGCUAAUUAGCCGCCGCUCUUUGUCCACUGGCCAA-CUU ....(((((....-((((((((..-..)))).)))))))))((((...(((.((..(((((.(--(---.((((....)))))))))))..)).)))...)))).-... ( -39.20) >DroAna_CAF1 20725 94 - 1 UCGUCG------CCUGUCGCUACC-CUUGGCUGAUUUUGCGUGGCAAUUGGCCAUUGAGCGAA--C---UGGCUAAUUAGCCGGCGCUCUUUGUCCACUUGGCAC-A-- .....(------(..(((((((..-..)))).)))...))(((.(((.(((.((..(((((..--(---(((((....)))))))))))..)).))).))).)))-.-- ( -38.00) >consensus UUGUCG______U_UGUCGCCACC_CUUGGCUGAUUUUGCGUGGCAAUUGGCCAUUGAGCGAG__C___UGGCUAAUUAGCCGGCGCUCUUUGUCCACUUGCCAA_CUU ..................((((.....))))..........((((((.(((.((..(((((........(((((....))))).)))))..)).))).))))))..... (-22.99 = -23.97 + 0.98)

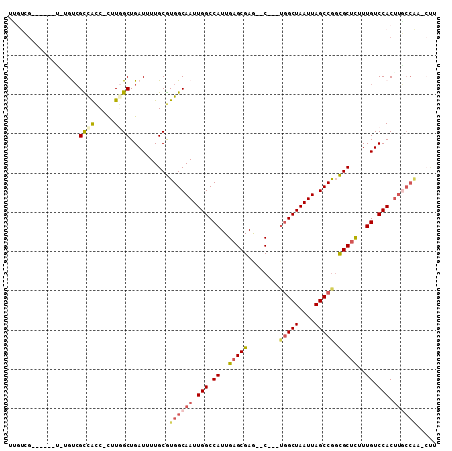
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:36:24 2006