
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 25,088,727 – 25,088,877 |
| Length | 150 |
| Max. P | 0.640292 |

| Location | 25,088,727 – 25,088,817 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 84.35 |
| Mean single sequence MFE | -26.97 |
| Consensus MFE | -20.95 |
| Energy contribution | -21.95 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588340 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25088727 90 - 27905053 AUGGGCUGGAUCGGCGGACACUCCUUUUAGGAGUCCGUA------AACCCUCUAAAU---AAAACAAG------------AAAACCUUU-UCCCGCUCUCAGUGGCAGCUCA .(((((((.....((((((..(((.....))))))))).------............---........------------.........-..((((.....))))))))))) ( -25.90) >DroSec_CAF1 17811 90 - 1 AUGGGCUGGAUCGGCGGACACUCCUUUUAGGAGUCCGUA------AACCCUCUAAAU---AAAACAAG------------GAAACCUUU-UCCCGCUCUCAGUGGCAGCUCA .(((((((.....((((((..(((.....))))))))).------............---.......(------------....)....-..((((.....))))))))))) ( -26.30) >DroSim_CAF1 17931 90 - 1 AUGGGCUGGAUCGGCGGACACUCCUUUUAGAAGUCCGUA------AAACCUCUAAAU---AAAACAAG------------GAAACCUUU-UCCCGCUCUCAGUGGCAGCUCA .(((((((.....((((((..((......)).)))))).------............---.......(------------....)....-..((((.....))))))))))) ( -22.80) >DroEre_CAF1 18192 93 - 1 AUGGGCUGGAUCGGCGGACACUCCUUUUAGGAGUCCGUA------AACCCUCUAAAU---AAAACAAAAAAAA---------AACCUUU-UCCCGCUCUCAGUGGCAGCUCA .(((((((.....((((((..(((.....))))))))).------............---.............---------.......-..((((.....))))))))))) ( -25.90) >DroYak_CAF1 18380 102 - 1 AUGGGCUGGAUCGGCGGACACUCCUUUUAGGAGACCGUA------AACCCUCUAAAU---AAAACAAAAAAAAAAAAGAGAAAACCUUU-UCCCGCUCUCAGUGGCAGCUCA .(((((((.....((((...((((.....)))).)))).------....((((....---................)))).........-..((((.....))))))))))) ( -24.85) >DroAna_CAF1 22759 103 - 1 AUGGGCUGGAUCGGCGGACACUCCUUUCAGCAGAGCGGAGUGCGAGGGCCUCAGGAUCCGAAAACAAAAGG---------GAAGCCUUUAUCCCGCUCCCAGUGGCAGCUCA .((((((((((((((...((((((.(((....))).)))))).....)))....))))).......(((((---------....)))))...((((.....)))).)))))) ( -36.10) >consensus AUGGGCUGGAUCGGCGGACACUCCUUUUAGGAGUCCGUA______AACCCUCUAAAU___AAAACAAA____________GAAACCUUU_UCCCGCUCUCAGUGGCAGCUCA .(((((((.....(((((..((((.....)))))))))......................................................((((.....))))))))))) (-20.95 = -21.95 + 1.00)



| Location | 25,088,781 – 25,088,877 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.67 |
| Mean single sequence MFE | -21.30 |
| Consensus MFE | -10.70 |
| Energy contribution | -10.15 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 25088781 96 + 27905053 ---GGACUCCUAAA-AGGAGUGUCCGCCGAUCCAGCCCAUAAUUAACUAUUCAAACAAUGGGCAC--------GCACAAAGGGAUCAGCAAUCAAAUACGCAUACA----CA-------- ---(((((((....-.)))..))))((.(((((.((((((.................)))))).(--------.......)))))).)).................----..-------- ( -26.03) >DroVir_CAF1 25195 106 + 1 ACGGCUUGACUCAA-AGGAGUGUACGCCGAUCCAACCCAUAAUUAACUAUUCAAACAAUAGAGAGAUGCGUGCCUACAA-AGGAUCGGCAAUCAAAUACGCAAACA----CA-------- ...((...((((..-..))))....((((((((.............(((((.....)))))........((....))..-.))))))))..........)).....----..-------- ( -23.00) >DroPse_CAF1 22488 102 + 1 ---GGU---CUGAAAAGGAGUGUCCGCUGAUCCAACCCAUAAUUAACUAUUCAAACAAUAGGCAU--------AUACAAAAGGAUCUGCAAUCAAAUACACACACA----CACACGGAAC ---..(---(((....((.....))((.(((((.............(((((.....)))))....--------........))))).)).................----....)))).. ( -16.31) >DroYak_CAF1 18446 96 + 1 ---GGUCUCCUAAA-AGGAGUGUCCGCCGAUCCAGCCCAUAAUUAACUAUUCAAACAAUGGGCAC--------GCACAAAGGGAUCAGCAAUCAAAUACGCAUACA----CA-------- ---((.((((....-.))))...))((.(((((.((((((.................)))))).(--------.......)))))).)).................----..-------- ( -25.53) >DroAna_CAF1 22826 96 + 1 ---GCUCUGCUGAA-AGGAGUGUCCGCCGAUCCAGCCCAUAAUUAACUAUUCAAACAAUAGGCAC--------ACACAAAGGGAUCAGCAAUCAAACACAUAUACA----UG-------- ---(((((......-.)))))....((.(((((.............(((((.....)))))....--------........))))).)).................----..-------- ( -16.51) >DroPer_CAF1 21976 105 + 1 ---GGU---CUGAA-AGGAGUGUCCGCUGAUCCAACCCAUAAUUAACUAUUCAAACAAUAGGCAU--------AUACAAAAGGAUCUGCAAUCAAAUACACACACACAGACACACGGAAC ---.((---(((..-.((.....))((.(((((.............(((((.....)))))....--------........))))).)).................)))))......... ( -20.41) >consensus ___GGUCU_CUGAA_AGGAGUGUCCGCCGAUCCAACCCAUAAUUAACUAUUCAAACAAUAGGCAC________ACACAAAAGGAUCAGCAAUCAAAUACACAUACA____CA________ ................((.....))((.(((((.............(((((.....))))).............(.....)))))).))............................... (-10.70 = -10.15 + -0.55)

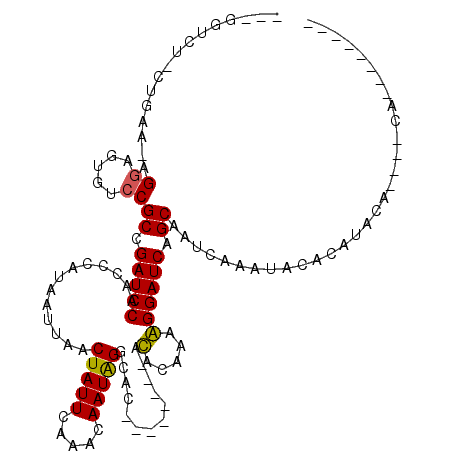

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:35:37 2006