
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,841,051 – 24,841,163 |
| Length | 112 |
| Max. P | 0.801546 |

| Location | 24,841,051 – 24,841,163 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 82.66 |
| Mean single sequence MFE | -28.42 |
| Consensus MFE | -17.19 |
| Energy contribution | -17.55 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.624058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24841051 112 + 27905053 UAA--GAUGACUUGCCAAAA--GUCUGGCAAAAAUAUUCCACUCCGCAAAUCAGGUCUCAAUUGGAAUAAUUCAAUUAAUAUUGAGUCAUGAAGUUGUCAUUGA-GUGCAGUUGCGG ...--......((((((...--...))))))............((((((.((((....(((((...((.(((((((....))))))).))..)))))...))))-......)))))) ( -25.70) >DroSec_CAF1 56834 114 + 1 UAACCGAUGACUUGCCAAAA--GUCUGGCAAAAAUAUUCCACUCCGCAAAUCAGGUCUCAAUUGGAAUAAUUCAAUUAAUAUUGAGUCAUGAAGUUGUCAUUGA-GUGCAGUUGCGG ...(((.((((((((((...--...))))))........(((((.((((.(((.(.((((((...(((......)))...)))))).).)))..))))....))-)))..))))))) ( -28.30) >DroSim_CAF1 60355 114 + 1 UAUCCGAUGACUUGCCAAAA--GUCUGGCAAAAAUAUUCCACUCCGCAAAUCAGGUCUCAAUUGGAAUAAUUCAAUUAAUAUUGAGUCAUGAAGUUGUCAUUGA-GUGCAGUUGCGG ...(((.((((((((((...--...))))))........(((((.((((.(((.(.((((((...(((......)))...)))))).).)))..))))....))-)))..))))))) ( -27.90) >DroEre_CAF1 54958 111 + 1 UAG--AAUGACUUGCCAAAA--GUCUGGCAA-AAUAUUCCACUCCGCAAAUCAAGCCUCAAUUGGAAUAAUUCAAUUAAUAAUGAGUUAUGAAGUUGUCAUCGG-CUGCAUUUGCGG ..(--((((..((((((...--...))))))-..)))))....((((((((..((((.(((((...((((((((........))))))))..))))).....))-))..)))))))) ( -35.70) >DroYak_CAF1 53073 112 + 1 UAA--GAUGACUUGCCAAAA--GUCUGGCAAAAAUAUUCCACUUCGCAAAUCAAAUCUCAAUUGGAAUAAUUCAAUUAAUAUUGAGCUAUGAAGUUGUCAUCAA-GUGCAUUUGCGC ...--((((((((((((...--...)))))).........((((((....((((.....(((((((....)))))))....))))....)))))).))))))..-((((....)))) ( -29.60) >DroAna_CAF1 51468 107 + 1 AGA--GAUGACUUGCCAAAAAAACUUUGCCGGAAUAUUCCACUCAAGAAAUCAAGCGUCGAUUGCAGUAAUUCAAUUAGUUUUGAGGUGGAA---UGUCAUCGGAAAACAGU----- ...--(((((((((.((((.....)))).)))...(((((((((((((......(((.....))).(.....)......)))))).))))))---)))))))..........----- ( -23.30) >consensus UAA__GAUGACUUGCCAAAA__GUCUGGCAAAAAUAUUCCACUCCGCAAAUCAAGUCUCAAUUGGAAUAAUUCAAUUAAUAUUGAGUCAUGAAGUUGUCAUCGA_GUGCAGUUGCGG .....((((((((((((........))))))...................(((.(.((((((...(((......)))...)))))).).)))....))))))............... (-17.19 = -17.55 + 0.36)

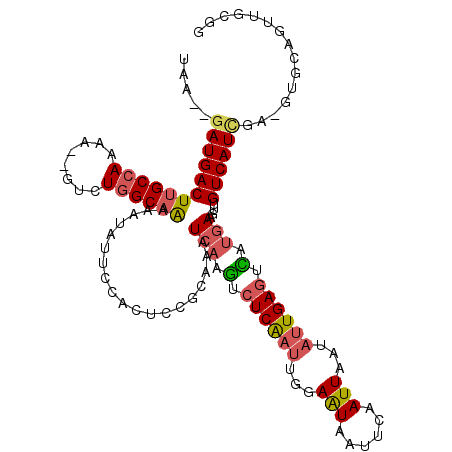

| Location | 24,841,051 – 24,841,163 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 82.66 |
| Mean single sequence MFE | -28.38 |
| Consensus MFE | -19.83 |
| Energy contribution | -20.88 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801546 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24841051 112 - 27905053 CCGCAACUGCAC-UCAAUGACAACUUCAUGACUCAAUAUUAAUUGAAUUAUUCCAAUUGAGACCUGAUUUGCGGAGUGGAAUAUUUUUGCCAGAC--UUUUGGCAAGUCAUC--UUA ..((....))..-...(((((.(((((..((.(((...(((((((........)))))))....))).))..))))).........(((((((..--..)))))))))))).--... ( -25.30) >DroSec_CAF1 56834 114 - 1 CCGCAACUGCAC-UCAAUGACAACUUCAUGACUCAAUAUUAAUUGAAUUAUUCCAAUUGAGACCUGAUUUGCGGAGUGGAAUAUUUUUGCCAGAC--UUUUGGCAAGUCAUCGGUUA ((((....))..-...(((((.(((((..((.(((...(((((((........)))))))....))).))..))))).........(((((((..--..)))))))))))).))... ( -26.60) >DroSim_CAF1 60355 114 - 1 CCGCAACUGCAC-UCAAUGACAACUUCAUGACUCAAUAUUAAUUGAAUUAUUCCAAUUGAGACCUGAUUUGCGGAGUGGAAUAUUUUUGCCAGAC--UUUUGGCAAGUCAUCGGAUA ((((....))..-...(((((.(((((..((.(((...(((((((........)))))))....))).))..))))).........(((((((..--..)))))))))))).))... ( -27.10) >DroEre_CAF1 54958 111 - 1 CCGCAAAUGCAG-CCGAUGACAACUUCAUAACUCAUUAUUAAUUGAAUUAUUCCAAUUGAGGCUUGAUUUGCGGAGUGGAAUAUU-UUGCCAGAC--UUUUGGCAAGUCAUU--CUA ((((((((..((-(((((((............))))).(((((((........)))))))))))..))))))))..((((((..(-(((((((..--..))))))))..)))--))) ( -36.50) >DroYak_CAF1 53073 112 - 1 GCGCAAAUGCAC-UUGAUGACAACUUCAUAGCUCAAUAUUAAUUGAAUUAUUCCAAUUGAGAUUUGAUUUGCGAAGUGGAAUAUUUUUGCCAGAC--UUUUGGCAAGUCAUC--UUA ..((....))..-..((((((.(((((.(((.((((..(((((((........)))))))...)))).))).))))).........(((((((..--..)))))))))))))--... ( -31.50) >DroAna_CAF1 51468 107 - 1 -----ACUGUUUUCCGAUGACA---UUCCACCUCAAAACUAAUUGAAUUACUGCAAUCGACGCUUGAUUUCUUGAGUGGAAUAUUCCGGCAAAGUUUUUUUGGCAAGUCAUC--UCU -----..........(((((((---((((((.((((........((((((..((.......)).)))))).))))))))))).....(.(((((...))))).)..))))))--... ( -23.30) >consensus CCGCAACUGCAC_UCAAUGACAACUUCAUAACUCAAUAUUAAUUGAAUUAUUCCAAUUGAGACCUGAUUUGCGGAGUGGAAUAUUUUUGCCAGAC__UUUUGGCAAGUCAUC__UUA ................(((((.(((((.(((.(((...(((((((........)))))))....))).))).))))).........((((((((....)))))))))))))...... (-19.83 = -20.88 + 1.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:32:20 2006