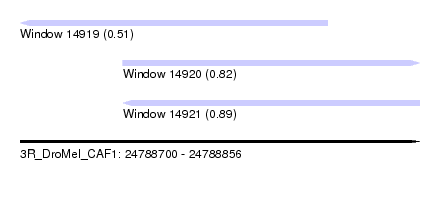

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,788,700 – 24,788,856 |
| Length | 156 |
| Max. P | 0.893912 |

| Location | 24,788,700 – 24,788,820 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.63 |
| Mean single sequence MFE | -32.60 |
| Consensus MFE | -32.35 |
| Energy contribution | -32.46 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.99 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24788700 120 - 27905053 CCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUAGCCAGCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUGAGAAUUUUAUCUGCUGUUCGGUAAUUCACAGCUCAGGCUC (((.(((((...(((((..((((......))))..)))))....))))).)))..(((..((((((.....))))))(((((.........((((....))))........)))))))). ( -34.63) >DroEre_CAF1 28880 116 - 1 CCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUA----GCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUGAGAAUUUUAUCUGCUGUUCGGUAAUUCACAGCUCAGGCUC (((.(((((...(((((..((((......))))..)))))----))))).)))..(((..((((((.....))))))(((((.........((((....))))........)))))))). ( -32.13) >DroYak_CAF1 20959 116 - 1 CCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGCA----GCCGUAUGGGUGGCAUUUUAACAAAGUGUUAAACUGAGAAUUUUAUCUGCUGUUCGGUAAUUCACAGCUCAGGCUC (((.(((((((.((((....)))).))(..(....)..).----))))).)))..(((..((((((.....))))))(((((.........((((....))))........)))))))). ( -31.03) >consensus CCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUA____GCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUGAGAAUUUUAUCUGCUGUUCGGUAAUUCACAGCUCAGGCUC (((.(((((...(((((..((((......))))..)))))....))))).)))..(((..((((((.....))))))(((((.........((((....))))........)))))))). (-32.35 = -32.46 + 0.11)



| Location | 24,788,740 – 24,788,856 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.26 |
| Mean single sequence MFE | -33.40 |
| Consensus MFE | -31.39 |
| Energy contribution | -31.17 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.819930 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24788740 116 + 27905053 CAGUUUAACACUUUGUUAAAAUGCUACCCAUACGGCUGGCUACGCUUACGUUCGUUUACGCUUACGUAUACGCCAUGUGGCGCGUCCGCCCAUAAAGUGUGCCAACAUUUGGCACC---- .......(((((((((......(((........))).(((.((((((((((.(((.((((....)))).)))..)))))).))))..))).)))))))))((((.....))))...---- ( -34.00) >DroEre_CAF1 28920 116 + 1 CAGUUUAACACUUUGUUAAAAUGCUACCCAUACGGC----UACGCUUACGUUCGUUUACGCUUACGUAUACGCCAUGUGGCGCGUCCGCCCAUAAAGUGUGCCAACACUUGGCACCCCGU .......(((((((((....(((.....)))..(((----.((((((((((.(((.((((....)))).)))..)))))).))))..))).)))))))))((((.....))))....... ( -31.70) >DroYak_CAF1 20999 116 + 1 CAGUUUAACACUUUGUUAAAAUGCCACCCAUACGGC----UGCGCUUACGUUCGUUUACGCUUACGUAUACGCCAUGUGGCGCGUCCGCCCAUAAAGUGUGCCAACAUUUGGCGCCCCGU .......(((((((((....(((.....))).(((.----((((((.((((.(((.((((....)))).)))..)))))))))).)))...)))))))))((((.....))))....... ( -34.50) >consensus CAGUUUAACACUUUGUUAAAAUGCUACCCAUACGGC____UACGCUUACGUUCGUUUACGCUUACGUAUACGCCAUGUGGCGCGUCCGCCCAUAAAGUGUGCCAACAUUUGGCACCCCGU .......(((((((((....((((...(((((.(((....((((....(((......)))....))))...))).))))).))))......)))))))))((((.....))))....... (-31.39 = -31.17 + -0.22)



| Location | 24,788,740 – 24,788,856 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.26 |
| Mean single sequence MFE | -40.13 |
| Consensus MFE | -36.70 |
| Energy contribution | -36.03 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.893912 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24788740 116 - 27905053 ----GGUGCCAAAUGUUGGCACACUUUAUGGGCGGACGCGCCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUAGCCAGCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUG ----.((((((.....))))))........(((((.((((((....))))).(((((..((((......))))..)))))))).)))....(..((((((((.....))))))))..).. ( -39.80) >DroEre_CAF1 28920 116 - 1 ACGGGGUGCCAAGUGUUGGCACACUUUAUGGGCGGACGCGCCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUA----GCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUG ((((.((((((.....))))))....((((((((....)))).))))((((.(((((...((....)).))))).)))).----.))))..(..((((((((.....))))))))..).. ( -40.40) >DroYak_CAF1 20999 116 - 1 ACGGGGCGCCAAAUGUUGGCACACUUUAUGGGCGGACGCGCCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGCA----GCCGUAUGGGUGGCAUUUUAACAAAGUGUUAAACUG .(((...((((..(((((((......((((((((....)))).))))((((.(((((...((....)).))))).)))).----)))).)))..))))..((((((.....))))))))) ( -40.20) >consensus ACGGGGUGCCAAAUGUUGGCACACUUUAUGGGCGGACGCGCCACAUGGCGUAUACGUAAGCGUAAACGAACGUAAGCGUA____GCCGUAUGGGUAGCAUUUUAACAAAGUGUUAAACUG ....(.((((......((((......((((((((....)))).))))((((.(((((...((....)).))))).)))).....))))....)))).)..((((((.....))))))... (-36.70 = -36.03 + -0.66)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:31:48 2006