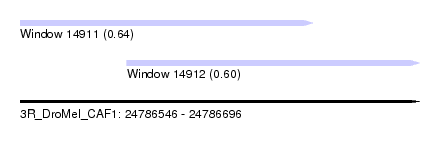
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,786,546 – 24,786,696 |
| Length | 150 |
| Max. P | 0.636039 |

| Location | 24,786,546 – 24,786,656 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.32 |
| Mean single sequence MFE | -34.13 |
| Consensus MFE | -26.49 |
| Energy contribution | -26.93 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636039 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24786546 110 + 27905053 AAUGACAAUCAAUCAUUCAGUUAGUUGCAGCAGCAAACUUCAGCAAUUUGGGGAC---------CCCCCUCGAAAACACCGCCCCUUUUUUUGGAGAGGGGGUCAUUAGGCC-CCCGCCC ((((((.............(((.((((.((.......)).))))..(((((((..---------...))))))))))....((((((((....)))))))))))))).(((.-...))). ( -31.90) >DroEre_CAF1 26724 109 + 1 AAUGACAAUCAAUCAUUCAGUUAGUUGCAGCAGCAAACUUCAGCAAUUUGGGGAU--------CCCCCCUCAAAAACACCGCCCA--UUUCGGGUGAGGGGGUCAUUAGGCC-CCCGCCC (((((.......))))).((((.((((...)))).))))...((..(((((((..--------....))))))).....(((((.--....))))).(((((((....))))-))))).. ( -35.50) >DroYak_CAF1 18626 120 + 1 AAUGACAAUCAAUCAUUCAGUUAGUUGCAGCAGCAAACUUCAGCAAUUUGGGGACUCCCCCCCCCCCCCUUAAAAACCCCGCCCACUUUUUGGGUGAGGGGGUCAUUAGGCCCCCCGCCC ..((.((((.(((......))).))))))((.((........)).....((((......)))).................)).........(((((.(((((((....)))))))))))) ( -35.00) >consensus AAUGACAAUCAAUCAUUCAGUUAGUUGCAGCAGCAAACUUCAGCAAUUUGGGGAC________CCCCCCUCAAAAACACCGCCCA_UUUUUGGGUGAGGGGGUCAUUAGGCC_CCCGCCC ((((((..................((((....)))).((((((....))))))............((((((.........((((.......)))))))))))))))).(((.....))). (-26.49 = -26.93 + 0.45)

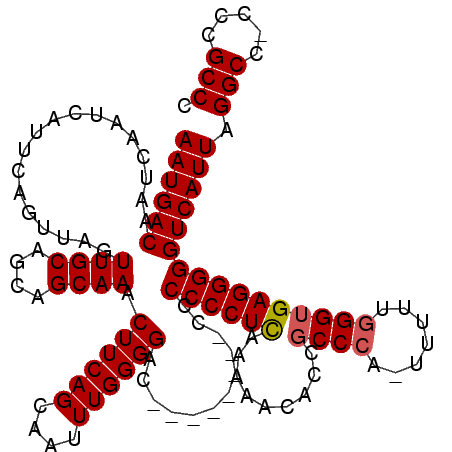
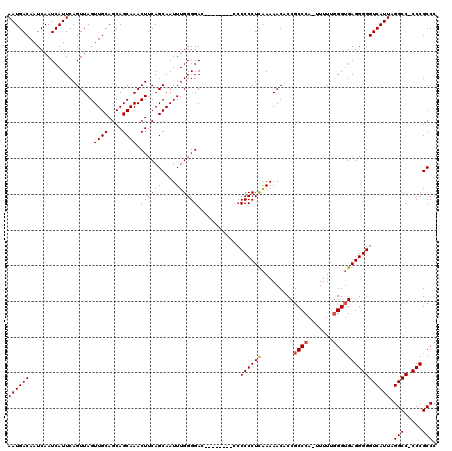
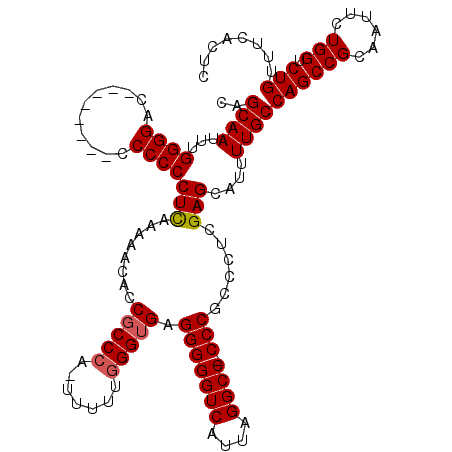
| Location | 24,786,586 – 24,786,696 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.32 |
| Mean single sequence MFE | -37.50 |
| Consensus MFE | -29.82 |
| Energy contribution | -30.27 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.604062 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24786586 110 + 27905053 CAGCAAUUUGGGGAC---------CCCCCUCGAAAACACCGCCCCUUUUUUUGGAGAGGGGGUCAUUAGGCC-CCCGCCCUCGAGCAUUUUGCCAGCCGCAAUUCUGGUCUGUUUCACUC ..(((((((((((((---------(((((((.((((............)))).))).)))))))....(((.-...)))))))))....))))((((((......))).)))........ ( -34.60) >DroEre_CAF1 26764 109 + 1 CAGCAAUUUGGGGAU--------CCCCCCUCAAAAACACCGCCCA--UUUCGGGUGAGGGGGUCAUUAGGCC-CCCGCCCUCGAGCAUUUUGCCAGCCGCAAUUCUGGUCUGUUUCACUC ..((((..(((((..--------..)))(((........(((((.--....))))).(((((((....))))-)))......)))))..))))((((((......))).)))........ ( -38.70) >DroYak_CAF1 18666 120 + 1 CAGCAAUUUGGGGACUCCCCCCCCCCCCCUUAAAAACCCCGCCCACUUUUUGGGUGAGGGGGUCAUUAGGCCCCCCGCCCUCGAGCAUUUUGCCAGCCGCAAUUCUGGUCUGUUUCACUC ..((((...((((......))))......................(((...(((((.(((((((....))))))))))))..)))....))))((((((......))).)))........ ( -39.20) >consensus CAGCAAUUUGGGGAC________CCCCCCUCAAAAACACCGCCCA_UUUUUGGGUGAGGGGGUCAUUAGGCC_CCCGCCCUCGAGCAUUUUGCCAGCCGCAAUUCUGGUCUGUUUCACUC ..((((...((((...........))))(((........(((((.......))))).(((((((....)))).)))......)))....))))((((((......))).)))........ (-29.82 = -30.27 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:31:41 2006