
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,771,015 – 24,771,149 |
| Length | 134 |
| Max. P | 0.879898 |

| Location | 24,771,015 – 24,771,119 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 80.66 |
| Mean single sequence MFE | -26.60 |
| Consensus MFE | -16.88 |
| Energy contribution | -17.77 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.797783 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24771015 104 + 27905053 GAGGGCGAGGGUGCCCUUUGCGGCCCAGAGAAAUUAAUAAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAACGUAUGCCGC---- (((((((....))))))).(((((...((....)).....((((.((.((((((........))))))(((((......)))))......)).))))..)))))---- ( -29.10) >DroPse_CAF1 23118 89 + 1 AAGGGCGU-------------------CAGAAAUUAAUAAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAGCGUGUGCCGCCCGA ..(((((.-------------------((...........((((.((.((((((........))))))(((((......)))))......)).)))).)).))))).. ( -21.80) >DroPer_CAF1 23420 89 + 1 AAGGGCGU-------------------CAGAAAUUAAUAAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCUGCGUCUGCCGCCCGA ..(((((.-------------------((((...(((((((.......)))))))(((((.......((((((......)))))).....))))).)))).))))).. ( -28.90) >consensus AAGGGCGU___________________CAGAAAUUAAUAAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAGCGUAUGCCGCCCGA ..(((((......................((..((((((((.......))))))))...))......((((((......))))))((((((...)))))).))))).. (-16.88 = -17.77 + 0.89)



| Location | 24,771,015 – 24,771,119 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 80.66 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -18.99 |
| Energy contribution | -19.43 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879898 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24771015 104 - 27905053 ----GCGGCAUACGUUGCGUAUGAGUGACAUUUUGAUCACUUUAUUUGACUGCUAAUAAAUGCGACGCUUAUUAAUUUCUCUGGGCCGCAAAGGGCACCCUCGCCCUC ----(((((...((((((((..(((((((.....).))))))((((........)))).))))))))((.............)))))))..(((((......))))). ( -30.82) >DroPse_CAF1 23118 89 - 1 UCGGGCGGCACACGCUGCGUAUGAGUGACAUUUUGAUCACUUUAUUUGACUGCUAAUAAAUGCGACGCUUAUUAAUUUCUG-------------------ACGCCCUU ..(((((((....)).((((..(((((((.....).))))))........(((........))))))).............-------------------.))))).. ( -22.90) >DroPer_CAF1 23420 89 - 1 UCGGGCGGCAGACGCAGCGUAUGAGUGACAUUUUGAUCACUUUAUUUGACUGCUAAUAAAUGCGACGCUUAUUAAUUUCUG-------------------ACGCCCUU ..(((((.((((.((((((...(((((((.....).))))))....)).))))((((((..((...))))))))...))))-------------------.))))).. ( -28.10) >consensus UCGGGCGGCACACGCUGCGUAUGAGUGACAUUUUGAUCACUUUAUUUGACUGCUAAUAAAUGCGACGCUUAUUAAUUUCUG___________________ACGCCCUU ..(((((((....)).((((..(((((((.....).))))))........(((........))))))).................................))))).. (-18.99 = -19.43 + 0.45)


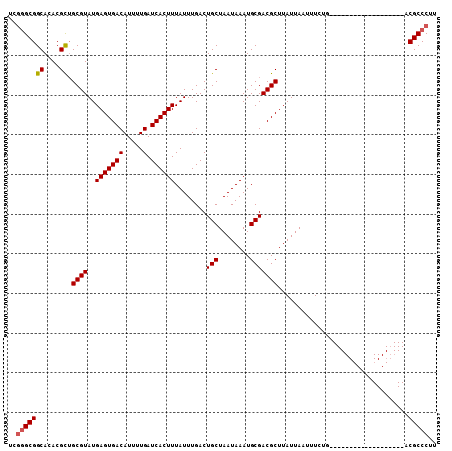
| Location | 24,771,053 – 24,771,149 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 94.33 |
| Mean single sequence MFE | -26.67 |
| Consensus MFE | -23.64 |
| Energy contribution | -24.53 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24771053 96 + 27905053 AAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAACGUAUGCCGC----CAGCUGAUGCUGUGGCAACAGUAAAAUAUU ..((((.((.((((((........))))))(((((......)))))......)).)))).((((((----.(((....)))))))))............. ( -25.50) >DroPse_CAF1 23137 100 + 1 AAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAGCGUGUGCCGCCCGACAGCUGAUGCUGUGGCAACGGUAAAAUAUU ....((((((..(((((((.(((......((((((......))))))((((((...))))))......))).))))))).)))))).............. ( -27.70) >DroPer_CAF1 23439 100 + 1 AAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCUGCGUCUGCCGCCCGACAGCUGAUGCUGUGGCAACGGUAAAAUAUU ....((((((..(((((((.(((......((((((......)))))).....((.((....)).))..))).))))))).)))))).............. ( -26.80) >consensus AAGCGUCGCAUUUAUUAGCAGUCAAAUAAAGUGAUCAAAAUGUCACUCAUACGCAGCGUAUGCCGCCCGACAGCUGAUGCUGUGGCAACGGUAAAAUAUU ....((((((..(((((((.(((......((((((......))))))((((((...))))))......))).))))))).)))))).............. (-23.64 = -24.53 + 0.89)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:30:59 2006