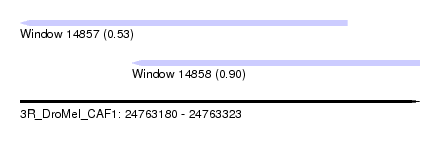
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,763,180 – 24,763,323 |
| Length | 143 |
| Max. P | 0.896983 |

| Location | 24,763,180 – 24,763,297 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.25 |
| Mean single sequence MFE | -23.67 |
| Consensus MFE | -21.35 |
| Energy contribution | -21.85 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.90 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
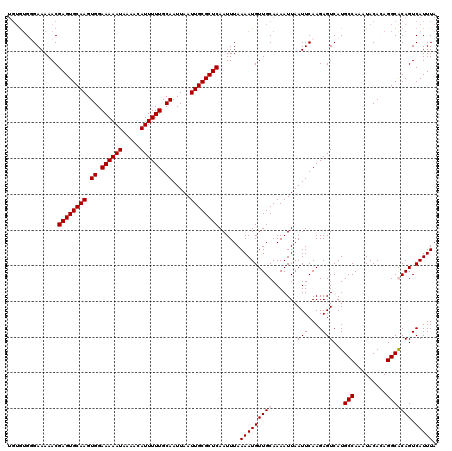
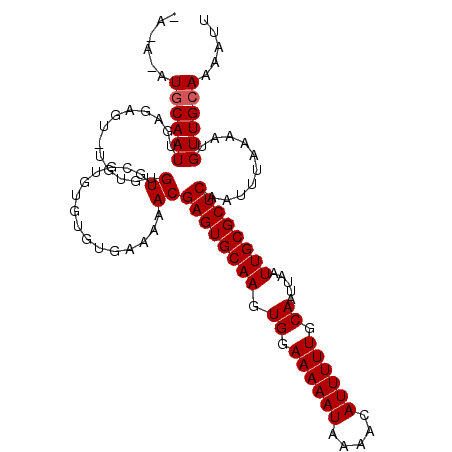
>3R_DroMel_CAF1 24763180 117 - 27905053 ---AUGUGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUUAAUUCAAGAGUCAUGCCAAAUACACAGGCGCAGUCAUUUA ---...........((((((((.((.((((((.....)))))).)).....))))))))......(((((((((.......(((....)))...(((.........))))))).))))). ( -24.80) >DroPse_CAF1 14608 120 - 1 UGUGUGGGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUUAAUUCAAGAGUCAUGCCAAAUACAGAGGCACAGUCAUUUA (((((.........((((((((.((.((((((.....)))))).)).....)))))))).........(((.(((..(((........)))..))).....)))...)))))........ ( -22.40) >DroWil_CAF1 12022 120 - 1 UGUGUGUGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUUAAUUCAAGAGUCAUGCCAAAUACACAGGCGCAGUCAUUUA (((((.((......((((((((.((.((((((.....)))))).)).....)))))))).............(((..(((........)))..)))........)).)))))........ ( -25.40) >DroPer_CAF1 14794 120 - 1 UGUGGGGGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUUAAUUCAAGAGUCAUGCCAAAUACAGAGGCACAGUCAUUUA ((..(.(.......((((((((.((.((((((.....)))))).)).....)))))))).........).)..))............((.(..((((.........)))).).))..... ( -22.09) >consensus UGUGUGGGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUUAAUUCAAGAGUCAUGCCAAAUACACAGGCACAGUCAUUUA ..............((((((((.((.((((((.....)))))).)).....))))))))......(((((((((.......(((....)))...(((.........))))))).))))). (-21.35 = -21.85 + 0.50)


| Location | 24,763,220 – 24,763,323 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 84.85 |
| Mean single sequence MFE | -24.49 |
| Consensus MFE | -18.19 |
| Energy contribution | -18.53 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.896983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
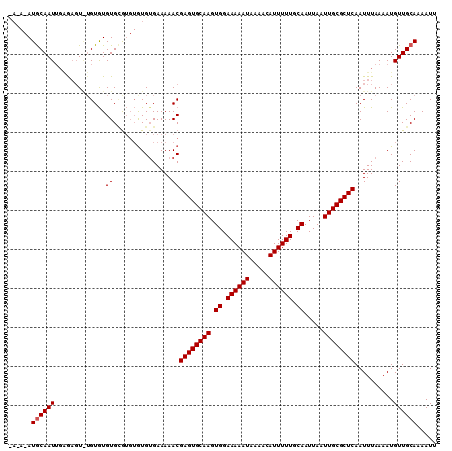
>3R_DroMel_CAF1 24763220 103 - 27905053 U----AUGCAAUGGAAAGUGUGUAUGUGCA---AUGUGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUU (----(((((.(....)...))))))((((---((((.......((((((((.((.((((((.....)))))).)).....))))))))........))))))))..... ( -25.76) >DroPse_CAF1 14648 110 - 1 CAGAAAUGCAAUUGGGUGUAUGUGUGUGGGUGUGUGGGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUU ......((((((...(..(((.(....).)))..).........((((((((.((.((((((.....)))))).)).....))))))))..........))))))..... ( -22.40) >DroWil_CAF1 12062 107 - 1 -AUAUAUACAAUUCUGAUU--GUGUGUGUGUGUGUGUGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUU -(((((((((((....)))--))))))))...(((((.....))((((((((.((.((((((.....)))))).)).....)))))))).............)))..... ( -25.30) >consensus _A_A_AUGCAAUUGAGAGU_UGUGUGUGCGUGUGUGUGAAAAACGAGUGCAAGUGGAAAAAUAAAACAUUUUUGCAAUUAAUUGCGCUCAAUUUAAAAUGUUGCAAAAUU ......((((((.............((...............))((((((((.((.((((((.....)))))).)).....))))))))..........))))))..... (-18.19 = -18.53 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:30:53 2006