
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,748,227 – 24,748,322 |
| Length | 95 |
| Max. P | 0.564224 |

| Location | 24,748,227 – 24,748,322 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -19.40 |
| Consensus MFE | -13.98 |
| Energy contribution | -13.57 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.564224 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24748227 95 + 27905053 ---UU-UUCAAAUGUGUAGAACCGUU--GCCGGCCUGUUUUGUUCGUUUUCGUU-UACAACCGAAUAUCGAAAGAUCAACC--CAUUUACACAUU----GCUACACAC------------ ---..-....((((((((((...(((--(.(((..(((......((....))..-.))).)))...(((....))))))).--..))))))))))----.........------------ ( -19.60) >DroSec_CAF1 2283 95 + 1 ---UU-UUGAAAUGUGUAGAGCCGUU--GCCGGCCUGUUUUGUUCGUUUUCGUU-UACAACCGAAUAUCGAAAGAUCAACC--CAUUUACACAUU----GCUACACAC------------ ---..-....((((((((((((((..--..)))).((.....((((..((((..-......))))...))))....))...--..))))))))))----.........------------ ( -21.70) >DroSim_CAF1 2282 94 + 1 ---UU-UUCAAAUGUGUAAAGCCGUU--GCCGGCCUGUUUUGUUCGUUUUCGUU-UACAACCGAAUAUCGAAAGAUCAACC--CAUUUACACA-U----GCUACACAC------------ ---..-.....(((((((((((((..--..)))).((.....((((..((((..-......))))...))))....))...--..))))))))-)----.........------------ ( -20.70) >DroEre_CAF1 2271 98 + 1 ---UUUUUCAAAUGUGUAGAACUGUC--GCCGGCCUGUUUUGUUCGUUUUCGUU-UACAACCGAAUAUCGAAAGAUCAACC--CAUUUACACAUU----GCUACACAC----------AC ---.........(((((((...(((.--...((..((.....((((..((((..-......))))...))))....))..)--)......)))..----.))))))).----------.. ( -17.90) >DroYak_CAF1 2276 107 + 1 ---UU-UUCAAAUGUGUAGAACUGUC--GCCGGCCUGUUUUGUUCGUUUUCGUU-UACAACCGAAUAUCGAAAGAUCAACC--CAUUCACACAUU----GCUACACACACACAAACACAC ---..-......(((((((...(((.--...((..((.....((((..((((..-......))))...))))....))..)--)......)))..----.)))))))............. ( -17.90) >DroPer_CAF1 2457 104 + 1 AUGUU-UUCAGAUGUGUAGAAUUAUCGCGCCAAUCCGUUUUGUUCGUUUUCGUUCUAUUGUCGAAUAUCGAAAACUCAACACACAUUUAUCCAUUACAUGCUAUA--------------- ((((.-...((((((((.((((....(((......)))...))))(((((((...((((....)))).))))))).....)))))))).......))))......--------------- ( -18.60) >consensus ___UU_UUCAAAUGUGUAGAACCGUC__GCCGGCCUGUUUUGUUCGUUUUCGUU_UACAACCGAAUAUCGAAAGAUCAACC__CAUUUACACAUU____GCUACACAC____________ ..........((((((((((.(((......))).........((((..((((.........))))...)))).............))))))))))......................... (-13.98 = -13.57 + -0.41)



| Location | 24,748,227 – 24,748,322 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -23.90 |
| Consensus MFE | -15.98 |
| Energy contribution | -15.90 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24748227 95 - 27905053 ------------GUGUGUAGC----AAUGUGUAAAUG--GGUUGAUCUUUCGAUAUUCGGUUGUA-AACGAAAACGAACAAAACAGGCCGGC--AACGGUUCUACACAUUUGAA-AA--- ------------((((((((.----..........((--..(((....((((...((((......-..))))..)))))))..))(((((..--..))))))))))))).....-..--- ( -24.70) >DroSec_CAF1 2283 95 - 1 ------------GUGUGUAGC----AAUGUGUAAAUG--GGUUGAUCUUUCGAUAUUCGGUUGUA-AACGAAAACGAACAAAACAGGCCGGC--AACGGCUCUACACAUUUCAA-AA--- ------------((((((((.----..........((--..(((....((((...((((......-..))))..)))))))..))(((((..--..))))))))))))).....-..--- ( -27.20) >DroSim_CAF1 2282 94 - 1 ------------GUGUGUAGC----A-UGUGUAAAUG--GGUUGAUCUUUCGAUAUUCGGUUGUA-AACGAAAACGAACAAAACAGGCCGGC--AACGGCUUUACACAUUUGAA-AA--- ------------.........----(-(((((((.((--..(((....((((...((((......-..))))..)))))))..))(((((..--..))))))))))))).....-..--- ( -26.00) >DroEre_CAF1 2271 98 - 1 GU----------GUGUGUAGC----AAUGUGUAAAUG--GGUUGAUCUUUCGAUAUUCGGUUGUA-AACGAAAACGAACAAAACAGGCCGGC--GACAGUUCUACACAUUUGAAAAA--- ..----------((((((((.----(((.(((...((--(.(((....((((...((((......-..))))..)))).....))).)))..--.))))))))))))))........--- ( -24.00) >DroYak_CAF1 2276 107 - 1 GUGUGUUUGUGUGUGUGUAGC----AAUGUGUGAAUG--GGUUGAUCUUUCGAUAUUCGGUUGUA-AACGAAAACGAACAAAACAGGCCGGC--GACAGUUCUACACAUUUGAA-AA--- ............((((((((.----(((.(((...((--(.(((....((((...((((......-..))))..)))).....))).)))..--.)))))))))))))).....-..--- ( -22.90) >DroPer_CAF1 2457 104 - 1 ---------------UAUAGCAUGUAAUGGAUAAAUGUGUGUUGAGUUUUCGAUAUUCGACAAUAGAACGAAAACGAACAAAACGGAUUGGCGCGAUAAUUCUACACAUCUGAA-AACAU ---------------..(((.((((..((((......(((((..((((((((...(((.......))))))))))...(.....)..)..))))).....)))).)))))))..-..... ( -18.60) >consensus ____________GUGUGUAGC____AAUGUGUAAAUG__GGUUGAUCUUUCGAUAUUCGGUUGUA_AACGAAAACGAACAAAACAGGCCGGC__AACAGUUCUACACAUUUGAA_AA___ ............((((((((....................(((.....((((...((((.........))))..))))...))).(((((......)))))))))))))........... (-15.98 = -15.90 + -0.08)

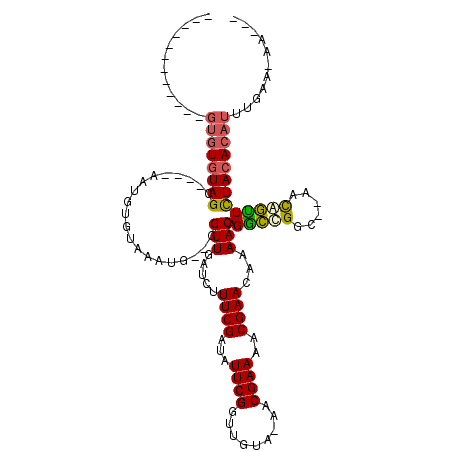

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:30:42 2006