
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,687,038 – 24,687,165 |
| Length | 127 |
| Max. P | 0.998906 |

| Location | 24,687,038 – 24,687,139 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 90.17 |
| Mean single sequence MFE | -21.06 |
| Consensus MFE | -15.54 |
| Energy contribution | -16.42 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.668879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24687038 101 + 27905053 UAUCUCUAUAUGGCCAGAGAACCAAACAU-ACCAUUCCCCAGAGACCGAUACCCAUAGAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCAUUGCUUCUCUA ..(((((........))))).........-..........(((((...........((((((((.....)))))))).......((((....))))))))). ( -18.80) >DroSec_CAF1 8410 101 + 1 UAUCUCUAUAUGGCCAGAGAGCCAAACAU-GCCAUUCCCCAGAGACCGAGACCCAUAGAUCUUGUUCUCCGAGGUCUCUACUAUAGCACCAUUGUUUCUCUA ...............((((((.(((...(-((.((.....(((((((((((..((.......)).))))...)))))))...)).)))...))).)))))). ( -22.60) >DroSim_CAF1 8369 101 + 1 UAUCUCUAUAUGGCCAGAGAGCCAAACAU-GCCAUUCCUCAGAGACCGAGACCCAUAGAUCUUGCUCUCCGAGGUCUCUACUAUAGCACCAUUGUUUCUCUA ...............((((((.(((...(-((.((.((((.((((.(((((........))))).)))).))))........)).)))...))).)))))). ( -24.30) >DroEre_CAF1 8482 102 + 1 UAUCUCUAUAUGACCAGAGAGCCAGACAUUACCAUUCCCCAACAAUCGAGACCCAUAGAUCUUGUUCUCUGUGGUCUCUACCAUAGCACCUUUGUUUCUCUA ...............(((((...........................((((((((.(((......))).)).))))))......((((....))))))))). ( -19.00) >DroYak_CAF1 8596 102 + 1 UAUCUCUAUAUGGCCAGAGAGUCAGAGAUUACCAUUCCCCAGACGACGAGACCCAUAAAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCUUUGUUUCUCUA ...............((((((.(((((..............((.(((((((........))))))).))...(((....))).......))))).)))))). ( -20.60) >consensus UAUCUCUAUAUGGCCAGAGAGCCAAACAU_ACCAUUCCCCAGAGACCGAGACCCAUAGAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCAUUGUUUCUCUA ..(((((........)))))....................(((((..(((((((..(((......)))..).))))))......((((....))))))))). (-15.54 = -16.42 + 0.88)


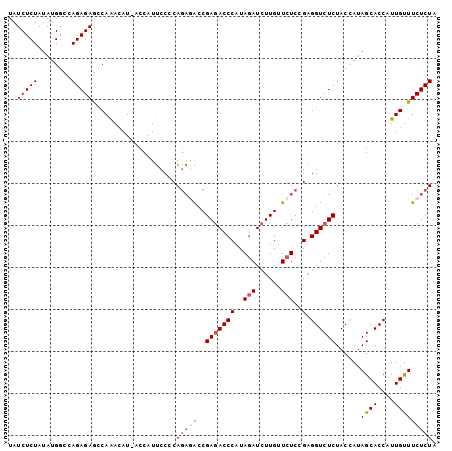
| Location | 24,687,038 – 24,687,139 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 90.17 |
| Mean single sequence MFE | -31.32 |
| Consensus MFE | -25.28 |
| Energy contribution | -25.72 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984636 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24687038 101 - 27905053 UAGAGAAGCAAUGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUCUAUGGGUAUCGGUCUCUGGGGAAUGGU-AUGUUUGGUUCUCUGGCCAUAUAGAGAUA (((((((.(((..(((((((........(((((((((.....(((....))).....))))))))).)))))-))..))).))))))).............. ( -32.00) >DroSec_CAF1 8410 101 - 1 UAGAGAAACAAUGGUGCUAUAGUAGAGACCUCGGAGAACAAGAUCUAUGGGUCUCGGUCUCUGGGGAAUGGC-AUGUUUGGCUCUCUGGCCAUAUAGAGAUA ((((((..(((..(((((((........(((((((((.(.(((((....))))).).))))))))).)))))-))..)))..)))))).............. ( -35.00) >DroSim_CAF1 8369 101 - 1 UAGAGAAACAAUGGUGCUAUAGUAGAGACCUCGGAGAGCAAGAUCUAUGGGUCUCGGUCUCUGAGGAAUGGC-AUGUUUGGCUCUCUGGCCAUAUAGAGAUA ((((((..(((..(((((((........(((((((((.(.(((((....))))).).))))))))).)))))-))..)))..)))))).............. ( -38.00) >DroEre_CAF1 8482 102 - 1 UAGAGAAACAAAGGUGCUAUGGUAGAGACCACAGAGAACAAGAUCUAUGGGUCUCGAUUGUUGGGGAAUGGUAAUGUCUGGCUCUCUGGUCAUAUAGAGAUA ((((((..((....((((((....((((((.((.(((......))).))))))))............)))))).....))..)))))).............. ( -24.09) >DroYak_CAF1 8596 102 - 1 UAGAGAAACAAAGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUUUAUGGGUCUCGUCGUCUGGGGAAUGGUAAUCUCUGACUCUCUGGCCAUAUAGAGAUA ((((((........(((....)))....(((((((..((.(((((....))))).))..)))))))........))))))..((((((......)))))).. ( -27.50) >consensus UAGAGAAACAAUGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUCUAUGGGUCUCGGUCUCUGGGGAAUGGU_AUGUUUGGCUCUCUGGCCAUAUAGAGAUA ((((((..(((....(((((........(((((((((.(.(((((....))))).).))))))))).))))).....)))..)))))).............. (-25.28 = -25.72 + 0.44)



| Location | 24,687,070 – 24,687,165 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 92.42 |
| Mean single sequence MFE | -22.68 |
| Consensus MFE | -17.52 |
| Energy contribution | -17.44 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.722667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24687070 95 + 27905053 AUUCCCCAGAGACCGAUACCCAUAGAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCAUUGCUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAAU ((((.((((..((((..(((...((((((((.....)))))))).......((((....))))........)))..))))..)....))).)))) ( -23.90) >DroSec_CAF1 8442 95 + 1 AUUCCCCAGAGACCGAGACCCAUAGAUCUUGUUCUCCGAGGUCUCUACUAUAGCACCAUUGUUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAAU ((((.((((..((((..(((...((((((((.....)))))))).......((((....))))........)))..))))..)....))).)))) ( -23.50) >DroSim_CAF1 8401 95 + 1 AUUCCUCAGAGACCGAGACCCAUAGAUCUUGCUCUCCGAGGUCUCUACUAUAGCACCAUUGUUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAAU ...((((.((((.(((((........))))).)))).))))((((((....((((....)))).....((((....)))).......)))))).. ( -24.20) >DroEre_CAF1 8515 95 + 1 AUUCCCCAACAAUCGAGACCCAUAGAUCUUGUUCUCUGUGGUCUCUACCAUAGCACCUUUGUUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAUU ..............((((((((.(((......))).)).))))))...........(((..(((....((((....))))....)))..)))... ( -23.10) >DroYak_CAF1 8629 95 + 1 AUUCCCCAGACGACGAGACCCAUAAAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCUUUGUUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAUU ............((((((........))))))((((((((((((.......)).))))).((((....((((....))))....))))))))).. ( -18.70) >consensus AUUCCCCAGAGACCGAGACCCAUAGAUCUUGUUCUCCGAGGUCUCUACCAUAGCACCAUUGUUUCUCUACUGGUUCCGGUUCCAAAAUGGAGAAU ..((.(((..(((((..(((...((((((((.....)))))))).......((((....))))........)))..)))))......))).)).. (-17.52 = -17.44 + -0.08)



| Location | 24,687,070 – 24,687,165 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.42 |
| Mean single sequence MFE | -28.96 |
| Consensus MFE | -26.38 |
| Energy contribution | -26.26 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.28 |
| SVM RNA-class probability | 0.998906 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24687070 95 - 27905053 AUUCUCCAUUUUGGAACCGGAACCAGUAGAGAAGCAAUGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUCUAUGGGUAUCGGUCUCUGGGGAAU ((((((((....(..(((((.(((.(((((..((((....))))..(((....))).............))))).))).)))))..))))))))) ( -29.70) >DroSec_CAF1 8442 95 - 1 AUUCUCCAUUUUGGAACCGGAACCAGUAGAGAAACAAUGGUGCUAUAGUAGAGACCUCGGAGAACAAGAUCUAUGGGUCUCGGUCUCUGGGGAAU ..((((.(((.(((.((((....).((......))...))).))).))).))))(((((((((.(.(((((....))))).).)))))))))... ( -29.90) >DroSim_CAF1 8401 95 - 1 AUUCUCCAUUUUGGAACCGGAACCAGUAGAGAAACAAUGGUGCUAUAGUAGAGACCUCGGAGAGCAAGAUCUAUGGGUCUCGGUCUCUGAGGAAU ..((((.(((.(((.((((....).((......))...))).))).))).))))(((((((((.(.(((((....))))).).)))))))))... ( -32.90) >DroEre_CAF1 8515 95 - 1 AAUCUCCAUUUUGGAACCGGAACCAGUAGAGAAACAAAGGUGCUAUGGUAGAGACCACAGAGAACAAGAUCUAUGGGUCUCGAUUGUUGGGGAAU ..((((((.((((....))))((((.(((....((....)).))))))).((((((.((.(((......))).))))))))......)))))).. ( -26.90) >DroYak_CAF1 8629 95 - 1 AAUCUCCAUUUUGGAACCGGAACCAGUAGAGAAACAAAGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUUUAUGGGUCUCGUCGUCUGGGGAAU ..((((.((..(((.((((....).((......))...))).)))..)).))))(((((((..((.(((((....))))).))..)))))))... ( -25.40) >consensus AUUCUCCAUUUUGGAACCGGAACCAGUAGAGAAACAAUGGUGCUAUGGUAGAGACCUCGGAGAACAAGAUCUAUGGGUCUCGGUCUCUGGGGAAU ..((((.(((.(((.((((....).((......))...))).))).))).))))(((((((((.(.(((((....))))).).)))))))))... (-26.38 = -26.26 + -0.12)


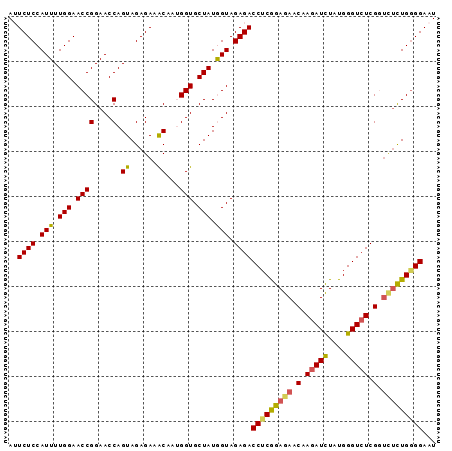
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:30:17 2006