
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,674,187 – 24,674,361 |
| Length | 174 |
| Max. P | 0.961213 |

| Location | 24,674,187 – 24,674,307 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.67 |
| Mean single sequence MFE | -29.76 |
| Consensus MFE | -28.12 |
| Energy contribution | -28.36 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961213 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24674187 120 - 27905053 UGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCUGCGACAGCGGAAUUCGCUGGAAAAGGCA .(((.((((......)))))))((((((((....))))))))..............((((....((((..(((....)))..)))).((((((...)))))).))))(((......))). ( -29.80) >DroSec_CAF1 15204 120 - 1 UGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCUGCGACAGCGGAAUUCGCUGGAAAAGGCA .(((.((((......)))))))((((((((....))))))))..............((((....((((..(((....)))..)))).((((((...)))))).))))(((......))). ( -29.80) >DroSim_CAF1 15245 120 - 1 UGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCUGCGACAGCGGAAUUCGCUGGAAAAGGCA .(((.((((......)))))))((((((((....))))))))..............((((....((((..(((....)))..)))).((((((...)))))).))))(((......))). ( -29.80) >DroEre_CAF1 16685 120 - 1 CGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCAGCGACAGCGGAAUUCGCUGGAAAAGGCA .....((((......)))).(((((((.....((((((((......))))))))..........((((..(((....)))..))))...)))))..(((((.....)))))......)). ( -29.60) >DroYak_CAF1 15619 120 - 1 CGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCAGCGACAGCGGAAUUCGCUGGAAAAAGCA .....((((......)))).(((((((.....((((((((......))))))))..........((((..(((....)))..))))...)))))..(((((.....)))))......)). ( -29.80) >consensus UGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACUGAAAUUGCCGCUGCGACAGCGGAAUUCGCUGGAAAAGGCA .(((.((((......)))))))((((((((....))))))))..............((((....((((..(((....)))..)))).((((((...)))))).))))(((......))). (-28.12 = -28.36 + 0.24)



| Location | 24,674,227 – 24,674,332 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 96.23 |
| Mean single sequence MFE | -26.90 |
| Consensus MFE | -26.36 |
| Energy contribution | -26.17 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.44 |
| SVM RNA-class probability | 0.954036 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24674227 105 - 27905053 GU--GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACU ((--((.....((((((.((((((..(((((.((((......))))))))).(((((....))))))))))).))))))......)))).................. ( -26.30) >DroSec_CAF1 15244 101 - 1 GU------GUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACU ..------...((((((.((((((..(((((.((((......))))))))).(((((....))))))))))).))))))............................ ( -25.70) >DroSim_CAF1 15285 105 - 1 GU--GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACU ((--((.....((((((.((((((..(((((.((((......))))))))).(((((....))))))))))).))))))......)))).................. ( -26.30) >DroEre_CAF1 16725 107 - 1 GUGUGUGUUUGUGUGCGGUUGGUUGUGCGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACU ....((((((.((((((.((((((..(((((.((((......))))))))).(((((....))))))))))).))))))....)))))).................. ( -29.30) >consensus GU__GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCGGCCAAAUGCAUAAAAUGAAUAUUAAAUUAAAGUUGAAACU ...........((((((.((((((..(((((.((((......))))))))).(((((....))))))))))).))))))............................ (-26.36 = -26.17 + -0.19)


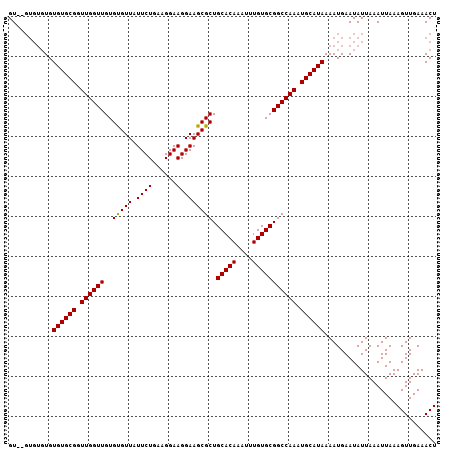
| Location | 24,674,267 – 24,674,361 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 95.79 |
| Mean single sequence MFE | -14.50 |
| Consensus MFE | -13.74 |
| Energy contribution | -13.74 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24674267 94 + 27905053 CGCACAAAUUUGUGCAGCGCUUCCUUCCUUCAGAAUAACACACAACCAACCGCACACACACAC--ACACACACGAGUGCGCAUAAAAUUGAUUCAU ....(((.(((((((.((((((..(((.....)))................(......)....--........))))))))))))).)))...... ( -13.80) >DroSec_CAF1 15284 90 + 1 CGCACAAAUUUGUGCAGCGCUUCCUUCCUUCAGAAUAACACACAACCAACCGCACACAC------ACACACACGAGUGCGCAUAAAAUUGAUUCAU ....(((.(((((((.((((((.....................................------........))))))))))))).)))...... ( -13.88) >DroSim_CAF1 15325 94 + 1 CGCACAAAUUUGUGCAGCGCUUCCUUCCUUCAGAAUAACACACAACCAACCGCACACACACAC--ACACACACGAGUGCGCAUAAAAUUGAUUCAU ....(((.(((((((.((((((..(((.....)))................(......)....--........))))))))))))).)))...... ( -13.80) >DroEre_CAF1 16765 96 + 1 CGCACAAAUUUGUGCAGCGCUUCCUUCCUUCAGAAUAACGCACAACCAACCGCACACAAACACACACACACACGAGUGCGCAUAAAAUUGAUUCAU ....(((.(((((((.((((((..(((.....)))....((..........))....................))))))))))))).)))...... ( -16.50) >consensus CGCACAAAUUUGUGCAGCGCUUCCUUCCUUCAGAAUAACACACAACCAACCGCACACACACAC__ACACACACGAGUGCGCAUAAAAUUGAUUCAU ....(((.(((((((.((((((...................................................))))))))))))).)))...... (-13.74 = -13.74 + 0.00)



| Location | 24,674,267 – 24,674,361 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 95.79 |
| Mean single sequence MFE | -29.62 |
| Consensus MFE | -25.66 |
| Energy contribution | -26.22 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.923543 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24674267 94 - 27905053 AUGAAUCAAUUUUAUGCGCACUCGUGUGUGU--GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCG ...(((((((..(((((((((.((......)--).)))))))))..)))))))..(((((.((((......))))))))).(((((....))))). ( -30.00) >DroSec_CAF1 15284 90 - 1 AUGAAUCAAUUUUAUGCGCACUCGUGUGUGU------GUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCG ...(((((((..(((((((((........))------)))))))..)))))))..(((((.((((......))))))))).(((((....))))). ( -29.70) >DroSim_CAF1 15325 94 - 1 AUGAAUCAAUUUUAUGCGCACUCGUGUGUGU--GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCG ...(((((((..(((((((((.((......)--).)))))))))..)))))))..(((((.((((......))))))))).(((((....))))). ( -30.00) >DroEre_CAF1 16765 96 - 1 AUGAAUCAAUUUUAUGCGCACUCGUGUGUGUGUGUGUUUGUGUGCGGUUGGUUGUGCGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCG ...(((((((....(((((((.((..(....)..))...))))))))))))))..(((((.((((......))))))))).(((((....))))). ( -28.80) >consensus AUGAAUCAAUUUUAUGCGCACUCGUGUGUGU__GUGUGUGUGUGCGGUUGGUUGUGUGUUAUUCUGAAGGAAGGAAGCGCUGCACAAAUUUGUGCG ...(((((((....(((((((.((..(......)..)).))))))))))))))..(((((.((((......))))))))).(((((....))))). (-25.66 = -26.22 + 0.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:30:03 2006