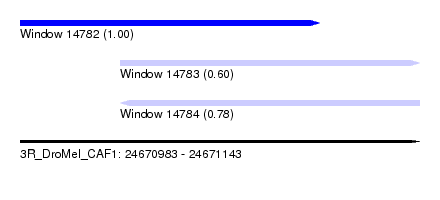
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,670,983 – 24,671,143 |
| Length | 160 |
| Max. P | 0.999173 |

| Location | 24,670,983 – 24,671,103 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -34.12 |
| Consensus MFE | -31.12 |
| Energy contribution | -32.40 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.41 |
| SVM RNA-class probability | 0.999173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24670983 120 + 27905053 UUUUGUUGUUGUUCUUCAAACAGCUGAGCGGCGGACGGAUAUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCGAAAAUUAAA (((((((((((((((((.........(((((((((......)))))))))(((((((..((....))))))))))))))))(((.............)))....))))))))))...... ( -39.32) >DroSec_CAF1 12063 120 + 1 UUUUGUUGUUGUUCUUCAAACAGCUGAGCGGCGGACGGAUAUUUGUUGUUGAUGCGCCCGGGAAGCAGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUUGCAAAAAUUAAA ((((((.((((((((((.....((((.((((((.(..((((.....))))..).))))..(....).)).)))))))))))(((.............)))....))).))))))...... ( -30.22) >DroSim_CAF1 12054 120 + 1 UUUUGUUGUUGUUCUUCAAACAGCUGAGCGGCGGACGGAUAUUUGUUGUUGCUCCGCCCGGGAAGCUGCGCAACGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCCAAAAUUAAA (((((((((((((((((...(((((..(.((((((..((((.....))))..)))))))....)))))......)))))))(((.............)))....))))).)))))..... ( -33.02) >DroEre_CAF1 13559 120 + 1 UUUUGUUGUUGUUUUACAAACAGCUGAGCGGCGGACGGAUAUUUGUUGAUGCUGCGCCCGGGAAGCUGCGCAGUGAAUAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAA ((((((((((..(((((..........(((((((((((....)))))..))))))((.(((....))).)).)))))....(((.............)))....))))))))))...... ( -28.12) >DroYak_CAF1 12194 120 + 1 UUUUGUUGUUGUUUUCCAAACAGCUGAGCGGCGGACGGAUAUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAA (((((((((((((((((.........(((((((((......)))))))))(((((((..((....))))))))))))))))(((.............)))....))))))))))...... ( -39.92) >consensus UUUUGUUGUUGUUCUUCAAACAGCUGAGCGGCGGACGGAUAUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAA (((((((((((((((((.........(((((((((......)))))))))(((((((..((....))))))))))))))))(((.............)))....))))))))))...... (-31.12 = -32.40 + 1.28)



| Location | 24,671,023 – 24,671,143 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.50 |
| Mean single sequence MFE | -30.93 |
| Consensus MFE | -20.38 |
| Energy contribution | -21.34 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597771 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24671023 120 + 27905053 AUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCGAAAAUUAAACGCGUUGCAAGUGCAGCAGAAAACAACGGUAAUAAUUCGC ..(((((((((((((((.......((.((((.(((.....)))..............(((........)))..........)))).))..))))))))....)))))))........... ( -32.90) >DroSec_CAF1 12103 120 + 1 AUUUGUUGUUGAUGCGCCCGGGAAGCAGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUUGCAAAAAUUAAACGUGUCGCGAGUGCAGCUGCAAACAACAGUAAUAAUUCGC ..(((((((((.(((((.((.((....(((..(((.....)))...............((((....))))..........))).)).)).)))))....).))))))))........... ( -29.70) >DroSim_CAF1 12094 120 + 1 AUUUGUUGUUGCUCCGCCCGGGAAGCUGCGCAACGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCCAAAAUUAAAAUCGUCGCAAGUGCAGCUGCAAACAACAGUAAUAAUUCGC ..(((((((((((((.....)))((((((((...(.....)((((.............((........))..............))))..)))))))))).))))))))........... ( -31.13) >DroEre_CAF1 13599 115 + 1 AUUUGUUGAUGCUGCGCCCGGGAAGCUGCGCAGUGAAUAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAACG-----CAAGUGCAGCAGCAAAAAAAAAUAAUAUUUCGC .(((((((..(((((((..((....))))))))).......((...............((........)).........((.-----...))))..)))))))................. ( -24.90) >DroYak_CAF1 12234 120 + 1 AUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAACGCGUCGCGAGUGCAGCAGCAAGCAACCAUAAUAAUUUGC ..(((((((((((((((.((.((...((((..(((.....)))...............((........))..........)))))).)).)))))))))).))))).............. ( -36.00) >consensus AUUUGUUGUUGCUGCGCCCGGGAAGCUGCGCAGCGGAGAACGCGAAAAAUAAUAAAACGCAAAAAAUAGCAAAAAUUAAACGCGUCGCAAGUGCAGCAGCAAACAACAGUAAUAAUUCGC ..((((((((...((.(....)..(((((((.(((.....)))...............((........))....................))))))).)).))))))))........... (-20.38 = -21.34 + 0.96)



| Location | 24,671,023 – 24,671,143 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.50 |
| Mean single sequence MFE | -30.38 |
| Consensus MFE | -22.10 |
| Energy contribution | -23.46 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.781679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24671023 120 - 27905053 GCGAAUUAUUACCGUUGUUUUCUGCUGCACUUGCAACGCGUUUAAUUUUCGCUAUUUUUUGCGUUUUAUUAUUUUUCGCGUUCUCCGCUGCGCAGCUUCCCGGGCGCAGCAACAACAAAU .............(((((....((((((.((((.((((((..((((...(((........)))....)))).....))))))....(((....)))....)))).))))))))))).... ( -32.70) >DroSec_CAF1 12103 120 - 1 GCGAAUUAUUACUGUUGUUUGCAGCUGCACUCGCGACACGUUUAAUUUUUGCAAUUUUUUGCGUUUUAUUAUUUUUCGCGUUCUCCGCUGCGCUGCUUCCCGGGCGCAUCAACAACAAAU ............(((((((.(((((.(....(((((......((((...(((((....)))))....))))....)))))....).)))))((.(((.....)))))...)))))))... ( -31.60) >DroSim_CAF1 12094 120 - 1 GCGAAUUAUUACUGUUGUUUGCAGCUGCACUUGCGACGAUUUUAAUUUUGGCUAUUUUUUGCGUUUUAUUAUUUUUCGCGUUCUCCGUUGCGCAGCUUCCCGGGCGGAGCAACAACAAAU ............(((((((.((((((((....((((.((...((((....((........)).....)))).)).))))((........))))))))(((.....))))))))))))... ( -28.40) >DroEre_CAF1 13599 115 - 1 GCGAAAUAUUAUUUUUUUUUGCUGCUGCACUUG-----CGUUUAAUUUUUGCUAUUUUUUGCGUUUUAUUAUUUUUCGCGUUAUUCACUGCGCAGCUUCCCGGGCGCAGCAUCAACAAAU ((((((..........))))))((((((.((((-----....((((...(((........)))....))))......((((........)))).......)))).))))))......... ( -25.10) >DroYak_CAF1 12234 120 - 1 GCAAAUUAUUAUGGUUGCUUGCUGCUGCACUCGCGACGCGUUUAAUUUUUGCUAUUUUUUGCGUUUUAUUAUUUUUCGCGUUCUCCGCUGCGCAGCUUCCCGGGCGCAGCAACAACAAAU .............((((.((((((((((.(..((((((((..((((...(((........)))....)))).....)))))....))).).)))(((.....)))))))))))))).... ( -34.10) >consensus GCGAAUUAUUACUGUUGUUUGCUGCUGCACUUGCGACGCGUUUAAUUUUUGCUAUUUUUUGCGUUUUAUUAUUUUUCGCGUUCUCCGCUGCGCAGCUUCCCGGGCGCAGCAACAACAAAU ((((((..........)))))).(((((.((((.((.((...((((...(((........)))....))))......((((........)))).)).)).)))).))))).......... (-22.10 = -23.46 + 1.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:29:48 2006