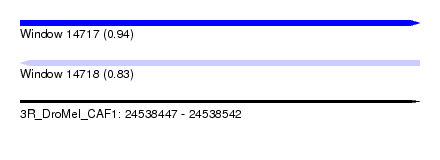
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,538,447 – 24,538,542 |
| Length | 95 |
| Max. P | 0.936148 |

| Location | 24,538,447 – 24,538,542 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -27.44 |
| Consensus MFE | -22.76 |
| Energy contribution | -23.76 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.936148 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24538447 95 + 27905053 GCCUAUGUAAAUUCACAGUUCACCGAUGGCUUAAAGCCAUAGAAGCACCUGCAAUCCCAGUGGAUUUCGCGAUAUGUGAUGGAAGUUAUAUAGGU ((((((((((.(((..(((((((.((((((.....)))))....((....)).....).)))))))((((.....))))..))).)))))))))) ( -28.70) >DroSec_CAF1 36702 95 + 1 GCCUAUGUAAAUUCACAGUUCACCGAUGGCUUAAAGCCGUAGAAGCACCUGCAAUCCCAUUGGAUUUCGCGGUAUGUGAUGGAAGUUAUAUAGGG .(((((((((..(((((....(((((((((.....))))).(((((....)).((((....))))))).)))).)))))......))))))))). ( -30.50) >DroSim_CAF1 38009 95 + 1 GCCUAUGUAAAUUCACAGUUCACCGAUGGCUUAAAGCCAUAGAAGCACCUGCAAUCCCAUUGGAUUUCGCGGUAUGUGAUGGAAGUUCUAUAGGG .((((((.((..(((((....(((((((((.....))))).(((((....)).((((....))))))).)))).)))))......)).)))))). ( -26.50) >DroEre_CAF1 41025 95 + 1 GCCCAUGUAAAUUCACAGUUCGCCGAUGGCUUAAAGCCAUAGAGCCACCUGCAAUUCCAUUGGAUUUCUCGGUAUGUGAUUGAAGGUAUAUAGGU (((.(((((..((((..((((....(((((.....))))).))))(((.(((...(((...))).......))).)))..))))..))))).))) ( -25.10) >DroYak_CAF1 37507 94 + 1 ACCCAUGUAAAUUCACAGUUCACCGAUGGCUUAAAGCCAUAGAGCCACCUGCAAU-CCAUUGGAUUUCGCGGCAUGUGAUGGUAGUUAUAUAGGU (((.((((((..(((((((((....(((((.....))))).))))...(((((((-((...)))))..))))..)))))......)))))).))) ( -26.40) >consensus GCCUAUGUAAAUUCACAGUUCACCGAUGGCUUAAAGCCAUAGAAGCACCUGCAAUCCCAUUGGAUUUCGCGGUAUGUGAUGGAAGUUAUAUAGGU .(((((((((..(((((....(((((((((.....)))))..........(.(((((....))))).).)))).)))))......))))))))). (-22.76 = -23.76 + 1.00)

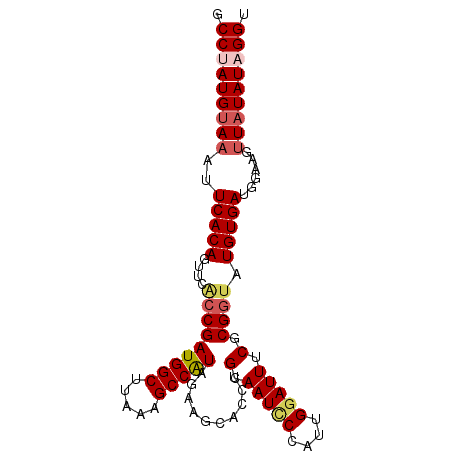

| Location | 24,538,447 – 24,538,542 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -26.28 |
| Consensus MFE | -20.04 |
| Energy contribution | -21.08 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830228 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24538447 95 - 27905053 ACCUAUAUAACUUCCAUCACAUAUCGCGAAAUCCACUGGGAUUGCAGGUGCUUCUAUGGCUUUAAGCCAUCGGUGAACUGUGAAUUUACAUAGGC .(((((.(((.(((((((((.((((((((..(((....))))))).))))...(.(((((.....))))).)))))..)).))).))).))))). ( -25.30) >DroSec_CAF1 36702 95 - 1 CCCUAUAUAACUUCCAUCACAUACCGCGAAAUCCAAUGGGAUUGCAGGUGCUUCUACGGCUUUAAGCCAUCGGUGAACUGUGAAUUUACAUAGGC .(((((.(((.(((((((((.((((((((..(((....))))))).)))).......(((.....)))....))))..)).))).))).))))). ( -26.20) >DroSim_CAF1 38009 95 - 1 CCCUAUAGAACUUCCAUCACAUACCGCGAAAUCCAAUGGGAUUGCAGGUGCUUCUAUGGCUUUAAGCCAUCGGUGAACUGUGAAUUUACAUAGGC .(((((..((.(((((((((.((((((((..(((....))))))).))))...(.(((((.....))))).)))))..)).))).))..))))). ( -26.40) >DroEre_CAF1 41025 95 - 1 ACCUAUAUACCUUCAAUCACAUACCGAGAAAUCCAAUGGAAUUGCAGGUGGCUCUAUGGCUUUAAGCCAUCGGCGAACUGUGAAUUUACAUGGGC .(((((.(((((.((((..(((...((....))..)))..)))).)))))(((..(((((.....))))).)))...............))))). ( -26.40) >DroYak_CAF1 37507 94 - 1 ACCUAUAUAACUACCAUCACAUGCCGCGAAAUCCAAUGG-AUUGCAGGUGGCUCUAUGGCUUUAAGCCAUCGGUGAACUGUGAAUUUACAUGGGU ((((((.(((.(((..((((..(((((..(((((...))-)))....))))).(.(((((.....))))).)))))...)))...))).)))))) ( -27.10) >consensus ACCUAUAUAACUUCCAUCACAUACCGCGAAAUCCAAUGGGAUUGCAGGUGCUUCUAUGGCUUUAAGCCAUCGGUGAACUGUGAAUUUACAUAGGC .(((((.(((.(((((((((.((((....(((((....)))))...))))...(.(((((.....))))).)))))..)).))).))).))))). (-20.04 = -21.08 + 1.04)

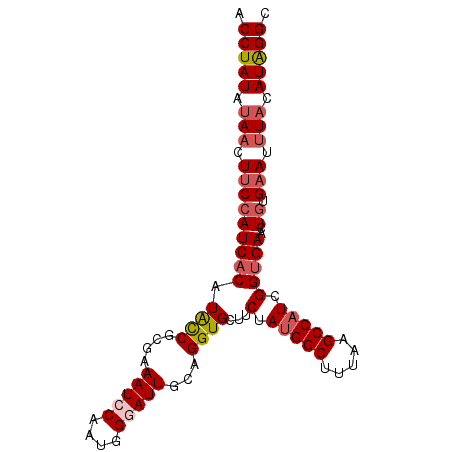
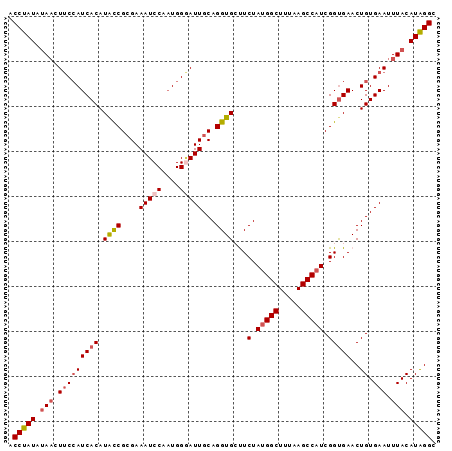
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:28:49 2006