
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,526,109 – 24,526,205 |
| Length | 96 |
| Max. P | 0.589701 |

| Location | 24,526,109 – 24,526,205 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 70.51 |
| Mean single sequence MFE | -41.63 |
| Consensus MFE | -16.56 |
| Energy contribution | -16.73 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.40 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589701 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24526109 96 - 27905053 CUGCUGCUGUUG---------CUGCUUC------UUGC------CGGCCAAGGACUCGGAAUCCUCGUGGCUAUCGCCGGGGUAGCCGCGUAUAAGCUGUGGCAGCAGGAGGCGCAG ((((.((....)---------).(((((------((((------..((((((((.......))))((((((((((.....)))))))))).........)))).))))))))))))) ( -44.90) >DroVir_CAF1 14301 102 - 1 CAGUUGUUGUUG---------CUGCUGU------UGUCCGGCAUCCACCACGGAGUCGCCAUCAUCCUGGUAAUCCGACGAGUAGCCGCGUAUAAGCUGCGGCAACAGCAGACGCAG .......(((..---------(((((((------((((((((........((((...((((......))))..))))(((........)))....)))).)))))))))))..))). ( -38.90) >DroPse_CAF1 30854 102 - 1 CUGGUGUUGUUG---------CUGCUUCUGCUGCUGCC------CGACCAGGGACUCGGCAUCGUCAUGAUUAUCCGCAGAAUAGCCGCGUAUUAGCUGCGGCAGCAGCAGACGCAA ...(((((..((---------((((((((((.((((((------(.....)))...))))(((.....))).....)))))...(((((((....)).))))))))))))))))).. ( -41.10) >DroMoj_CAF1 32689 111 - 1 GUGUUGUUGUUGUUGCCGCUGCUGCUGC------GGUCCAGCAGCCGCCAAGGAGUCGCCAUCGUCCGGGCAAUCAGCAGCGUAGCCGCGUAUCAGCUGCGGCAACAGCAGACGCAG (((((..(((((((((((((((((((((------......))))).(((..(((..((....))))).)))....))))))(((((.........)))))))))))))))))))).. ( -53.80) >DroAna_CAF1 25263 102 - 1 CUGCUGCUUUUGUUU------CUGCUUC---UGCUGAC------CGGCCAAGGACUCUGCAUACUCCUGGUCAUCAACGUGGGAGCCUCGUAUUAGCAGCGGCAAAAGAAGACGGAG (((((.(((((((..------(((((..---((((((.------.(((((.(((..........)))))))).)))..(.((...)).))))..)))))..))))))).)).))).. ( -32.40) >DroPer_CAF1 10346 102 - 1 CUGGUGUUGUUG---------CUGUUUCUGCUGCUGCC------CGACCAGGGACUCGGCAUCGUCAUGAUUAUCCGCAGAAUAGCCGCGUAUUAGCUGCGGCAACAGCAGACGCAA ...(((((..((---------((((((((((.((((((------(.....)))...))))(((.....))).....)))))...(((((((....)).))))))))))))))))).. ( -38.70) >consensus CUGCUGUUGUUG_________CUGCUUC______UGCC______CGACCAAGGACUCGGCAUCGUCCUGGUUAUCCGCAGAGUAGCCGCGUAUUAGCUGCGGCAACAGCAGACGCAG ...((((((((..................................(((((.((...........)).)))))............(((((((....)).)))))))))))))...... (-16.56 = -16.73 + 0.17)

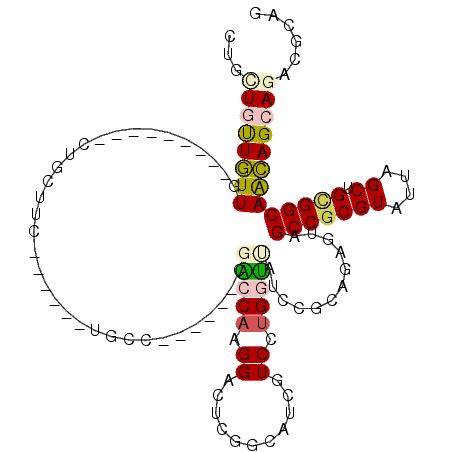
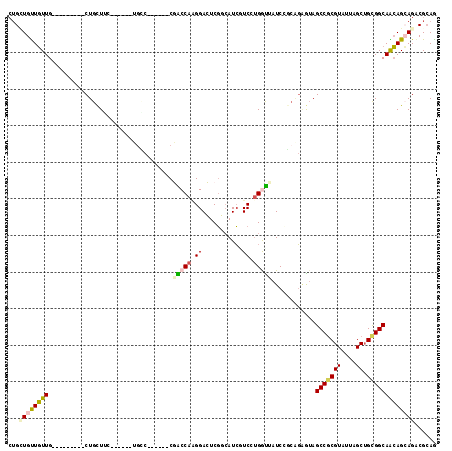
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:28:43 2006