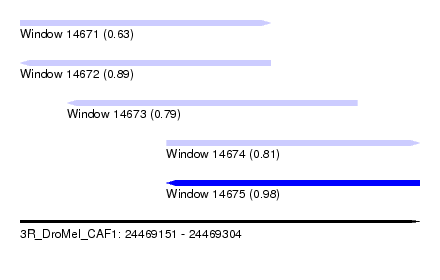

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,469,151 – 24,469,304 |
| Length | 153 |
| Max. P | 0.976775 |

| Location | 24,469,151 – 24,469,247 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -32.16 |
| Consensus MFE | -23.38 |
| Energy contribution | -23.70 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.627549 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24469151 96 + 27905053 --------GUCCAGUGGGCAAGAACAACUCGUUUUGCGGCUGAGAUUG--UGUUUGUUGCCCCAUCUAUAUGGCUGCAUGGCUGGAGGUAUAGCUGUAGCUCUAAC --------.....(.(((((((((((.(((((......)).)))....--))))).))))))).....((.(((((((..((((......))))))))))).)).. ( -27.70) >DroSec_CAF1 12293 94 + 1 ----------CCAGUAGGCAACAACAACUCGUGUUGUGGCCGAGAUUG--UGUUUGUUGCCUCAUCUAUAUGGCUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAAC ----------...(.((((((((((((((((.((....)))))).)))--)...))))))))).....((((((((((..((((......)))))))))))))).. ( -35.80) >DroSim_CAF1 24102 96 + 1 --------GUCCAGUGGGCAACAACAACUCGUGUUGUGGCUGAGAUUG--UGUUUGUUGCCUCAUCUAUAUGGGUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAAC --------.((((((((((((((((((((((.((....)))))).)))--)...))))))))(((((....)))))....))))))..((((((....)))))).. ( -34.10) >DroEre_CAF1 15733 99 + 1 -------CUACCAGUGGGCAACAACAAGUUGUGUUGUGGCUGAGAUUGAGUGUUUGUUGCCCCAUCUGAAUGGCUGCAUGGCCGAAGCUAUAGCUUUAGCUAUAAC -------....(((.(((((((((((....((.((......)).))....)).)))))))))...)))..(((((....)))))....((((((....)))))).. ( -30.40) >DroYak_CAF1 21935 104 + 1 ACGUCUACGUCCAGUGGGCAACAACAAGUUGUGUUGUGGCCGAGAUUG--UGUUUGUUGGCCCAUCUGAAUGGCUGCAUGGCAGAAGCUAGAGCUGUUGCUAUAAC ..((((((.....)))))).......(((..(..((.((((((.(...--....).)))))))).....)..)))..(((((((.(((....))).)))))))... ( -32.80) >consensus ________GUCCAGUGGGCAACAACAACUCGUGUUGUGGCUGAGAUUG__UGUUUGUUGCCCCAUCUAUAUGGCUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAAC ...........(((((((((((((((....((.(((....))).))....)).)))))))))(((....))))))).(((((((.(((....))).)))))))... (-23.38 = -23.70 + 0.32)



| Location | 24,469,151 – 24,469,247 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -19.16 |
| Energy contribution | -20.20 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.889020 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24469151 96 - 27905053 GUUAGAGCUACAGCUAUACCUCCAGCCAUGCAGCCAUAUAGAUGGGGCAACAAACA--CAAUCUCAGCCGCAAAACGAGUUGUUCUUGCCCACUGGAC-------- ((...(((....)))..)).(((((.........(((....)))((((((.....(--(((.(((...........)))))))..)))))).))))).-------- ( -21.40) >DroSec_CAF1 12293 94 - 1 GUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUAGAUGAGGCAACAAACA--CAAUCUCGGCCACAACACGAGUUGUUGUUGCCUACUGG---------- ((((((((....)))))))).((((.........(((....)))((((((((...(--(((.((((.........)))))))))))))))).))))---------- ( -32.10) >DroSim_CAF1 24102 96 - 1 GUUAUAGCUACAGCUAUAGCUCCAGCCAUGCACCCAUAUAGAUGAGGCAACAAACA--CAAUCUCAGCCACAACACGAGUUGUUGUUGCCCACUGGAC-------- ((((((((....))))))))(((((..(((....))).....((.(((((((...(--(((.(((...........))))))))))))))))))))).-------- ( -29.40) >DroEre_CAF1 15733 99 - 1 GUUAUAGCUAAAGCUAUAGCUUCGGCCAUGCAGCCAUUCAGAUGGGGCAACAAACACUCAAUCUCAGCCACAACACAACUUGUUGUUGCCCACUGGUAG------- ((((((((....))))))))...(((......)))..((((...(((((((((.((........................))))))))))).))))...------- ( -29.76) >DroYak_CAF1 21935 104 - 1 GUUAUAGCAACAGCUCUAGCUUCUGCCAUGCAGCCAUUCAGAUGGGCCAACAAACA--CAAUCUCGGCCACAACACAACUUGUUGUUGCCCACUGGACGUAGACGU ((((.(((....))).)))).(((((..(.(((((((....))))...........--.......(((.((((((.....)))))).)))..))).).)))))... ( -25.70) >consensus GUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUAGAUGGGGCAACAAACA__CAAUCUCAGCCACAACACGAGUUGUUGUUGCCCACUGGAC________ ((((((((....))))))))(((((.........(((....)))((((((((.................((((......)))))))))))).)))))......... (-19.16 = -20.20 + 1.04)


| Location | 24,469,169 – 24,469,280 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 90.07 |
| Mean single sequence MFE | -25.50 |
| Consensus MFE | -18.56 |
| Energy contribution | -20.04 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.789078 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
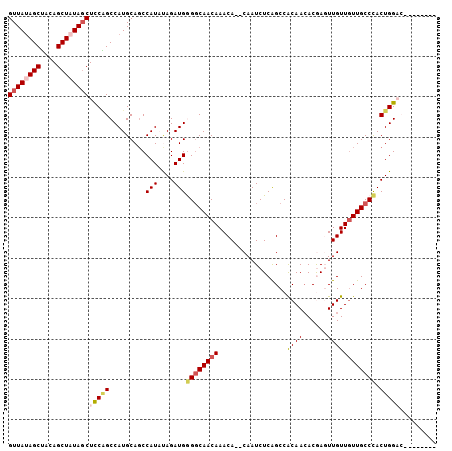
>3R_DroMel_CAF1 24469169 111 - 27905053 AAACAAACCAAAAGAUCACACCGUUUUGGCUUGGUUAGAGCUACAGCUAUACCUCCAGCCAUGCAGCCAUAUAGAUGGGGCAACAAACA--CAAUCUCAGCCGCAAAACGAGU ...................((((((((((((.(((...(((....)))..)))...)))))(((..((((....))))(((........--........))))))))))).)) ( -26.29) >DroSec_CAF1 12309 111 - 1 AAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUAGAUGAGGCAACAAACA--CAAUCUCGGCCACAACACGAGU ......................(((.(((((.(((((((((....)))))))))...(((...((..........)).)))........--.......))))).)))...... ( -26.40) >DroSim_CAF1 24120 111 - 1 AAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCACCCAUAUAGAUGAGGCAACAAACA--CAAUCUCAGCCACAACACGAGU ......................(((.(((((((((((((((....)))))))..))))...(((.(.(((....))).)))).......--........)))).)))...... ( -22.90) >DroEre_CAF1 15752 113 - 1 AAACAAACCAGAAAAUCAGACCGUUUUGGCCUGGUUAUAGCUAAAGCUAUAGCUUCGGCCAUGCAGCCAUUCAGAUGGGGCAACAAACACUCAAUCUCAGCCACAACACAACU ..................((..(((((((((.(((((((((....)))))))))..)))))(((..((((....)))).)))..))))..))..................... ( -28.60) >DroYak_CAF1 21961 111 - 1 AAACAAACCAGAAAAUCAGACCGUUUUGGCCUGGUUAUAGCAACAGCUCUAGCUUCUGCCAUGCAGCCAUUCAGAUGGGCCAACAAACA--CAAUCUCGGCCACAACACAACU ......................(((.(((((.(((((.(((....))).))))).((((...))))((((....))))...........--.......))))).)))...... ( -23.30) >consensus AAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUAGAUGGGGCAACAAACA__CAAUCUCAGCCACAACACGAGU ......................(((.(((((.(((((((((....)))))))))...(((......((((....))))))).................))))).)))...... (-18.56 = -20.04 + 1.48)


| Location | 24,469,207 – 24,469,304 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 88.24 |
| Mean single sequence MFE | -31.64 |
| Consensus MFE | -23.00 |
| Energy contribution | -23.48 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807648 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
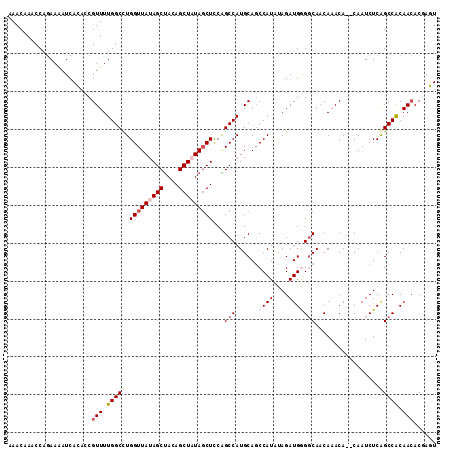
>3R_DroMel_CAF1 24469207 97 + 27905053 UAUAUGGCUGCAUGGCUGGAGGUAUAGCUGUAGCUCUAACCAAGCCAAAACGGUGUGAUCUUUUGGUUUGUUUGCGGAAAGGUGCAAUAGCAGUAGC------ ......(((((.(((((...(((..(((....)))...))).)))))......(((.(((((((.((......)).))))))))))...)))))...------ ( -29.90) >DroSec_CAF1 12347 97 + 1 UAUAUGGCUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAACCAGGCCAAAACGGUGUGAUUUUCUGGUUUGUUUGCGGAAAGGUGCAAUAGCAGUAGC------ ......((((((((((((.(((....))).)))))))(((((((((.....)))........))))))...(((((......)))))..)))))...------ ( -30.70) >DroSim_CAF1 24158 97 + 1 UAUAUGGGUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAACCAGGCCAAAACGGUGUGAUUUUCUGGUUUGUUUGUGGAAAGGUGCACUAGCAGUAGC------ ..(((.((((((((((((.(((....))).))))))).(((...(((((((((...(.....)....)))))).)))...))))))))....)))..------ ( -28.90) >DroEre_CAF1 15792 103 + 1 UGAAUGGCUGCAUGGCCGAAGCUAUAGCUUUAGCUAUAACCAGGCCAAAACGGUCUGAUUUUCUGGUUUGUUUGCGGAAAGGUGCACUAGCAAUAGCAAUAGC ....(((((....)))))..(((((.(((...((((..((((((((.....)))))...((((((.........)))))))))....))))...))).))))) ( -31.70) >DroYak_CAF1 21999 103 + 1 UGAAUGGCUGCAUGGCAGAAGCUAGAGCUGUUGCUAUAACCAGGCCAAAACGGUCUGAUUUUCUGGUUUGUUUGCGGAAAGGUGCAAUAGCAAUAGCAAUAGC .......((((...))))..((((..(((((((((((.((((((((.....)))))...((((((.........)))))))))...)))))))))))..)))) ( -37.00) >consensus UAUAUGGCUGCAUGGCUGGAGCUAUAGCUGUAGCUAUAACCAGGCCAAAACGGUGUGAUUUUCUGGUUUGUUUGCGGAAAGGUGCAAUAGCAGUAGC______ ......((((((((((((.(((....))).)))))))(((((((((.....)))........))))))((((((((......))))..)))))))))...... (-23.00 = -23.48 + 0.48)


| Location | 24,469,207 – 24,469,304 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 88.24 |
| Mean single sequence MFE | -25.20 |
| Consensus MFE | -19.10 |
| Energy contribution | -19.90 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.78 |
| SVM RNA-class probability | 0.976775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
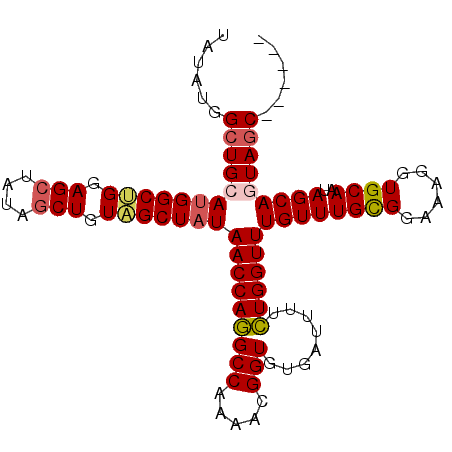
>3R_DroMel_CAF1 24469207 97 - 27905053 ------GCUACUGCUAUUGCACCUUUCCGCAAACAAACCAAAAGAUCACACCGUUUUGGCUUGGUUAGAGCUACAGCUAUACCUCCAGCCAUGCAGCCAUAUA ------....((((..((((........))))........................(((((.(((...(((....)))..)))...))))).))))....... ( -21.70) >DroSec_CAF1 12347 97 - 1 ------GCUACUGCUAUUGCACCUUUCCGCAAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUA ------....((((..((((........))))........................((((..(((((((((....)))))))))...)))).))))....... ( -24.60) >DroSim_CAF1 24158 97 - 1 ------GCUACUGCUAGUGCACCUUUCCACAAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCACCCAUAUA ------((....))..(((((..((((.............))))............((((..(((((((((....)))))))))...)))))))))....... ( -22.92) >DroEre_CAF1 15792 103 - 1 GCUAUUGCUAUUGCUAGUGCACCUUUCCGCAAACAAACCAGAAAAUCAGACCGUUUUGGCCUGGUUAUAGCUAAAGCUAUAGCUUCGGCCAUGCAGCCAUUCA (((.((((...(((....))).......))))....................((..(((((.(((((((((....)))))))))..))))).)))))...... ( -27.80) >DroYak_CAF1 21999 103 - 1 GCUAUUGCUAUUGCUAUUGCACCUUUCCGCAAACAAACCAGAAAAUCAGACCGUUUUGGCCUGGUUAUAGCAACAGCUCUAGCUUCUGCCAUGCAGCCAUUCA ((((..(((.((((((((((........))))...((((((....(((((....))))).)))))).)))))).)))..))))..((((...))))....... ( -29.00) >consensus ______GCUACUGCUAUUGCACCUUUCCGCAAACAAACCAGAAAAUCACACCGUUUUGGCCUGGUUAUAGCUACAGCUAUAGCUCCAGCCAUGCAGCCAUAUA ......(((..(((....)))...............................((..((((..(((((((((....)))))))))...)))).)))))...... (-19.10 = -19.90 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:28:12 2006