
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,447,063 – 24,447,197 |
| Length | 134 |
| Max. P | 0.904834 |

| Location | 24,447,063 – 24,447,166 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 79.69 |
| Mean single sequence MFE | -14.82 |
| Consensus MFE | -9.32 |
| Energy contribution | -10.72 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.788223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24447063 103 - 27905053 UACAC-A-AAA-A-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGCCCUCCAGCACCACCACUACUUCAUUUUCUCCUUUUCCCUUUAAUUUCGCUGCCAGCCCACAGC .....-.-...-.-((((.(((((((..........))))))).))))...........................................((((........)))) ( -15.80) >DroSim_CAF1 21104 103 - 1 UACAC-A-AAA-A-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGCCCUCCAGCACCACCACUACUUCAUUUUCUUCGUUUCCCUUUAAUUUCGCUGCCAGCCCACAGC .....-.-...-.-((((.(((((((..........))))))).))))...........................................((((........)))) ( -15.80) >DroEre_CAF1 20979 104 - 1 UACAC-A-AAAAA-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAGCACCACCACUACUUAAUUUUCUCCUUCUUCCUUCAAUUUUGCUGCCAGCCCACAGA .....-.-.....-((((.(((((((..........))))))).))))(((((.....................................)))))............ ( -15.09) >DroYak_CAF1 22002 105 - 1 UACAC-A-AAAAAGGAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAGCACCGCCACUUCUUAAUUUUCUCCUUUUGCCUUCAAUUCUGCUGCCAGCCCACAGC ....(-(-(((..(((((.(((((((..........))))))).))))).((...))..................)))))...........((((........)))) ( -19.90) >DroAna_CAF1 21307 82 - 1 CCCAACAAAAA-A-GAGGCCAUGAAUUAAAUUUCGUAUUCAUCUGC-----CA-CCCCACU-------UUUCCACU-----CAUCCAUUUUCGUG-----CCACAGC ...........-.-..(((.((((((..........))))))..))-----).-.......-------........-----..............-----....... ( -7.50) >consensus UACAC_A_AAA_A_GAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAGCACCACCACUACUUAAUUUUCUCCUUUUCCCUUCAAUUUUGCUGCCAGCCCACAGC ..............((((.(((((((..........))))))).))))...........................................((((........)))) ( -9.32 = -10.72 + 1.40)



| Location | 24,447,103 – 24,447,197 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 70.55 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -5.78 |
| Energy contribution | -7.45 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.27 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.904834 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24447103 94 - 27905053 ---UCGAAUGGGGAGCACAA-G-UUGC-CUUUAAGUUUACAC-A-AAA-A-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGCCCUCCAG--CACC------ACCACUACUUC---A ---..((((((((.((....-(-(.((-......))..))..-.-...-.-((((.(((((((..........))))))).))))..)--).))------.)))....)))---. ( -21.10) >DroSim_CAF1 21144 94 - 1 ---UCGAAUGGGGAGCACAA-G-UUCC-CUUAAAGUUUACAC-A-AAA-A-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGCCCUCCAG--CACC------ACCACUACUUC---A ---..(((.(((((((....-)-))))-))............-.-...-.-((((.(((((((..........))))))).))))...--....------........)))---. ( -23.20) >DroEre_CAF1 21019 96 - 1 ---UCGAAUGGGGAACACAA-G-UUCCCCUUUAAGUUUACAC-A-AAAAA-GAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAG--CACC------ACCACUACUUA---A ---......(((((((....-)-)))))).((((((......-.-.....-((((.(((((((..........))))))).))))...--....------......)))))---) ( -27.01) >DroWil_CAF1 21860 109 - 1 AAAAGGGAUGGGAAAGAAGAGGGUAUAACUUUAAGUUUCCGU-A---A-A-GUUGCCAUGAAUAAAAUUUCGAAUUCUUUUGCCACAAACCACUGUACAAACGAAUACUUCUGAA ....((..((((..(((((((((((..((((((........)-)---)-)-)))))).((((......))))..))))))).)).))..))........................ ( -19.20) >DroYak_CAF1 22042 97 - 1 ---UCGAAUGGGCAACACAA-G-UUCCCCUUUAAGUUUACAC-A-AAAAAGGAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAG--CACC------GCCACUUCUUA---A ---..(((..((((((....-)-)).................-.-.....(((((.(((((((..........))))))).)))))..--....------)))..)))...---. ( -22.70) >DroAna_CAF1 21336 81 - 1 -------AAAGGCAGCGUAG-G-AUAUCCUUAAAGUGCCCAACAAAAA-A-GAGGCCAUGAAUUAAAUUUCGUAUUCAUCUGC-------CA-C------CCCACU--------- -------...((((.(..((-(-....)))....))))).........-.-..(((.((((((..........))))))..))-------).-.------......--------- ( -15.70) >consensus ___UCGAAUGGGGAACACAA_G_UUCCCCUUUAAGUUUACAC_A_AAA_A_GAGGCCAUGAAUUAAAUUUUGUAUUCAUGUCCUCCAG__CACC______ACCACUACUUC___A ...................................................((((.(((((((..........))))))).)))).............................. ( -5.78 = -7.45 + 1.67)

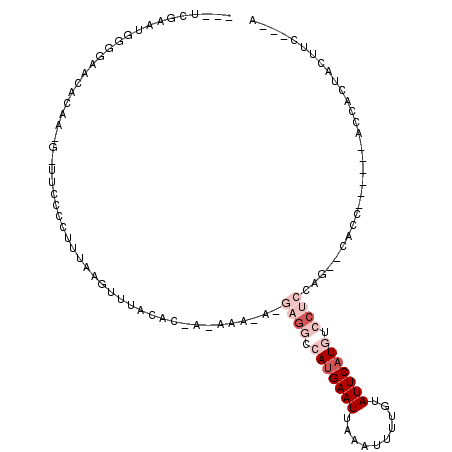

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:28:05 2006