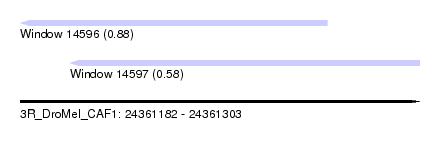
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,361,182 – 24,361,303 |
| Length | 121 |
| Max. P | 0.884552 |

| Location | 24,361,182 – 24,361,275 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 92.47 |
| Mean single sequence MFE | -23.50 |
| Consensus MFE | -20.43 |
| Energy contribution | -21.48 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.884552 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
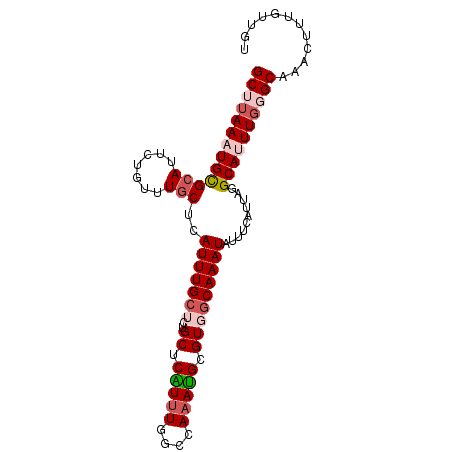
>3R_DroMel_CAF1 24361182 93 - 27905053 GCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCACUGCUCAUUUGGUCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGGGCAAACUUUGUUGU (((((((((((((.......)))..((((((...((.(((((....))))).)).))))))..........))))))))))............ ( -26.10) >DroGri_CAF1 166538 92 - 1 GCUCAACUGUGCAUUCUGUUUGCUCAUUUGCUCCGCUCGUUUGCUCAAACGCGUGACAAAUAUUACAUUAAGCAUUUAUGCAAACUUUGUUG- ((.(....).)).....((((((..(..((((.(((.(((((....))))).))).....((......))))))..)..)))))).......- ( -18.30) >DroSim_CAF1 191544 93 - 1 GCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCACUGCUCGUUUGGUCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGGGCAAACUUUGUUGU (((((((((((((.......)))..((((((...((.(((((....))))).)).))))))..........))))))))))............ ( -25.70) >DroEre_CAF1 195439 93 - 1 GCUUAAAUGCGCAUUCUGUUUACUCAUUUGCUCUGCUCAUUUGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGGGCAAACUUUGUUGU ((((((((((((.....))......(((((((..((.(((((....))))).)))))))))..........))))))))))............ ( -23.10) >DroYak_CAF1 189334 93 - 1 GCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCUCUGCUCAUUUGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGGGCAAACUUUGUUGU (((((((((((((.......)))..(((((((..((.(((((....))))).)))))))))..........))))))))))............ ( -26.70) >DroAna_CAF1 185281 92 - 1 GCUUAACUGUGCAUUCUGUUUGCUCAUUUGCUCUGCUCGU-UGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGUGCAAACUUUGUUGU ((........)).....((((((.((..((((.((...((-(.((((......)))).)))....))...))))..)).))))))........ ( -21.10) >consensus GCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCUCUGCUCAUUUGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUGGGCAAACUUUGUUGU (((((((((((((.......)))..(((((((..((.(((((....))))).)))))))))..........))))))))))............ (-20.43 = -21.48 + 1.06)


| Location | 24,361,197 – 24,361,303 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 78.69 |
| Mean single sequence MFE | -25.67 |
| Consensus MFE | -13.41 |
| Energy contribution | -13.50 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.583515 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24361197 106 - 27905053 ----UUUCUCAGGCCCAGUCUCGGUGAAC-UGGGCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCACUGCUCAUUUGGUCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUG ----......((((((((((.....).))-)))))))(((((((((.......)))..((((((...((.(((((....))))).)).))))))..........)))))). ( -32.40) >DroVir_CAF1 169066 107 - 1 ----UAUGCCUGUCUCUUUCUAUCUGUGCUAGAGCUUUACUGUGCAUUCUGUUUGCUCAUUUGCUUUGCUCGUUUGUUCAAACGCGUGACAAAUAUUUCAUUAGGCAUUUA ----.(((((((.............(.((.(((((.....((.(((.......))).))...))))))).)((((((.((......)))))))).......)))))))... ( -23.00) >DroGri_CAF1 166552 107 - 1 ----UUUCUUCGUCUCUUUCUAUCUGAGCUUGAGCUCAACUGUGCAUUCUGUUUGCUCAUUUGCUCCGCUCGUUUGCUCAAACGCGUGACAAAUAUUACAUUAAGCAUUUA ----.......(((...........((((.(((((..(((.(......).))).)))))...)))).((.(((((....))))).)))))..................... ( -21.20) >DroYak_CAF1 189349 106 - 1 ----UUUCUCGCGCCCAGUCUCGGUAAAC-UGGGCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCUCUGCUCAUUUGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUG ----........(((((((........))-))))).((((((((((.......)))..(((((((..((.(((((....))))).)))))))))..........))))))) ( -31.20) >DroMoj_CAF1 145624 101 - 1 ----UUUAUUUGUCCCUGUCUAACU------GAGCUUCAAUGCGAAUUCAGUUUGCUCAUUUACUUUGCUCGUUUGCUCAAACGCGUGACAAAUAUUUCAUUAGGCAUUUA ----..((((((((...((..((((------(((.(((.....)))))))))).))...........((.(((((....))))).))))))))))................ ( -20.30) >DroAna_CAF1 185296 106 - 1 UCUGUCUCUGCGACUCUGUCU---UUGAC-UGCGCUUAACUGUGCAUUCUGUUUGCUCAUUUGCUCUGCUCGU-UGGCCAAAUGCGUGGCAAAUAUUUCAUUAGGCAUUUG ...(((.....)))..(((((---.(((.-(((((......)))))...((((((((((((((..(.......-.)..))))))...))))))))..)))..))))).... ( -25.90) >consensus ____UUUCUCCGUCCCUGUCUAGCUGAAC_UGAGCUUAAAUGCGCAUUCUGUUUGCUCAUUUGCUCUGCUCGUUUGGUCAAACGCGUGACAAAUAUUUCAUUAGGCAUUUA .................................((((((..(.(((.......))).)(((((..((((.(((((....))))).)))))))))......))))))..... (-13.41 = -13.50 + 0.09)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:27:04 2006