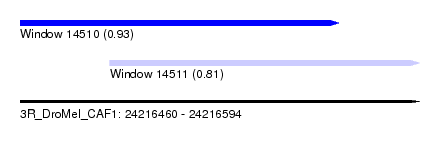
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 24,216,460 – 24,216,594 |
| Length | 134 |
| Max. P | 0.931656 |

| Location | 24,216,460 – 24,216,567 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.82 |
| Mean single sequence MFE | -34.82 |
| Consensus MFE | -22.42 |
| Energy contribution | -22.48 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.931656 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24216460 107 + 27905053 GA--G----CCCGCCU-UCUCCAAUCUC---GUUAGCCAGCCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCG---CACACU ..--.----...((((-...........---(((.((((((.....)))))))))((....)).)))).((((..((((((((.((((........))))..)))))))))---)))... ( -36.40) >DroPse_CAF1 39699 112 + 1 AAUGA----UCCGCAU-UCUCCGUCCACAGUGUUGGCCAAACGCUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCU---GUGUCU .....----.......-........(((((.((((((((((((((....)))..((((.((....)).))))...)))))....((((........))))...))))))))---)))... ( -34.20) >DroGri_CAF1 12437 101 + 1 ---------GCCGCGCUGUUGCCACUAA---GCUGGGCGGACAUU-GUUGGCAGCACUUCCGGCAGUCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCC---AAC--- ---------...(((((((((((...((---(.((.((.(((...-))).))..)))))..))))).))))))....((((((.((((........))))..))))))...---...--- ( -35.50) >DroSec_CAF1 39101 102 + 1 -----------CGCCU-UCUCCAACCUC---GUUAGCCAGCCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCG---CUCACC -----------.((((-...........---(((.((((((.....)))))))))((....)).)))).(((...((((((((.((((........))))..)))))))).---..))). ( -34.00) >DroAna_CAF1 36811 113 + 1 GA--AUUUUACCGCCC-UCUUCACUCUC---GUUAGCCCGCCGG-AGUUGGAAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACUGGCCCCCGCC ..--........(((.-.(((((((...---....(((.(.(((-((.((....)).))))).).))))))).))).((((((.((((........))))..))))))..)))....... ( -34.41) >DroPer_CAF1 39734 112 + 1 AAUGA----UCCGCGU-UCUCCGUCCACAGUGUUGGCCAAACGCUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCU---GUGUCU .....----...(((.-....))).(((((.((((((((((((((....)))..((((.((....)).))))...)))))....((((........))))...))))))))---)))... ( -34.40) >consensus _A__A____UCCGCCU_UCUCCAACCAC___GUUAGCCAGACGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCG___CUCACU ............((.................(((.(((((.......))))))))(((.((....)).)))))....((((((.((((........))))..))))))............ (-22.42 = -22.48 + 0.06)



| Location | 24,216,490 – 24,216,594 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 83.97 |
| Mean single sequence MFE | -33.37 |
| Consensus MFE | -25.20 |
| Energy contribution | -25.28 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.811947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 24216490 104 + 27905053 CCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCGCACACUCCUGCGU-------CCUUCGUCCUGCCUCG-GUCU .((..(((.(....))))..))((((((((..(..((((((((.((((........))))..))))))))(((......)))..-------.....)..)))))).)-)... ( -36.70) >DroPse_CAF1 39734 105 + 1 ACGCUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCUGUGUCUCCAUUCC-------ACCCAGCACUGCAUCCACUCU (((..(((.(....))))..)))...(((((((.((.((((((.((((........))))..)))))).))(((.........)-------))...)))))))......... ( -32.40) >DroSec_CAF1 39126 104 + 1 CCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCGCUCACCCCUUCGU-------CCUUCGUCCUGCCUCG-GUCU .((..(((.(....))))..))((((((((..(..((((((((.((((........))))..))))))))............(.-------....))..)))))).)-)... ( -32.30) >DroEre_CAF1 43892 110 + 1 CCGUAAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCGCUCACUCCUUCGUCCUGCGUCCUUCGUCCUGCU-CG-GGCU .((.((((.(....))))).))..(((.((((...((((((((.((((........))))..))))))))...)))))))..((((.(((..........))).-.)-))). ( -35.00) >DroYak_CAF1 41648 103 + 1 CCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAACUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCGCUCACUCUUUCGU-------CCUUCGUCCUGCC-UG-UUCU .((..(((.(....))))..)).(((((((..(....((((((.((((........))))..))))))(((..........)))-------.....)..)))))-))-.... ( -31.40) >DroPer_CAF1 39769 105 + 1 ACGCUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAACCUGUGUCUCCAUUCC-------ACCCAGCACUGCAUCCACUCU (((..(((.(....))))..)))...(((((((.((.((((((.((((........))))..)))))).))(((.........)-------))...)))))))......... ( -32.40) >consensus CCGUUAGUUGGCAACACUUCCGUCAGGCAGUGCAAGUUUGACACGCGCAUCAACACGCGCUUUGUCAAGCGCUCACUCCUUCGU_______CCUUCGUCCUGCCUCG_GUCU .((..(((.(....))))..))...(((((.((..((((((((.((((........))))..))))))))...(........).............)).)))))........ (-25.20 = -25.28 + 0.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:25:48 2006