
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,906,881 – 23,906,975 |
| Length | 94 |
| Max. P | 0.999743 |

| Location | 23,906,881 – 23,906,975 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 92.35 |
| Mean single sequence MFE | -27.14 |
| Consensus MFE | -22.66 |
| Energy contribution | -22.90 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814910 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23906881 94 + 27905053 UCUGGCCAUU-UGGUCCCAUAAUGGCCAUUGGGCUGACCAGACUUCAAUUAUUUGUGUCUGGCCCACGACGCCCACGGAUUGAAAAUGGGAAAUU ..((((((((-((....)).)))))))).(((((....(((((..(((....))).)))))))))).....((((...........))))..... ( -31.20) >DroSec_CAF1 107442 89 + 1 UCUGGCCAUU-UGGUCCCAUAAUGGCCAUUGGGCUGACCAGACUCCAAUUAUUUGUGUCUGGCCCACGACGCCCACGGAUUGAAAAUGGG----- ..((((((((-((....)).)))))))).(((((((.((((((..(((....))).))))))..))....)))))...............----- ( -30.40) >DroSim_CAF1 110135 91 + 1 UCUGGCCAUU-UGGUCCCAUAAUAGCCAUUGGGCUGACCAGACUCCAAUUAUUUGUGUCUGGCCCACGACGCCCACGGAUUGAAAAUGGGAA--- ....(((...-.)))(((((.....((..(((((((.((((((..(((....))).))))))..))....))))).)).......)))))..--- ( -27.30) >DroEre_CAF1 111496 94 + 1 UCUGGCUAUU-UGGUCCCAUAAUGGCCAUUGGACUGACCAGACUCCAAUUAUUUGUGUCUGCCCCACGACGCCCACGGAUUGAAAAUGGAAAAUU ..((((((((-((....)).)))))))).........(((.....(((((....(((((........))))).....)))))....)))...... ( -22.30) >DroYak_CAF1 115109 95 + 1 UUUGGCUAUUUUGGUCCCAUAAUGGCCAUUGCACUGACCAGACUCCAAUUAUUUGUGUCUGCCCCACGACGCCCACGGAUUGAAAAUGCGAAAUU ..((((((((.((....)).))))))))(((((((....))....(((((....(((((........))))).....)))))....))))).... ( -24.50) >consensus UCUGGCCAUU_UGGUCCCAUAAUGGCCAUUGGGCUGACCAGACUCCAAUUAUUUGUGUCUGGCCCACGACGCCCACGGAUUGAAAAUGGGAAAUU ..((((((((.((....)).)))))))).........(((.....(((((....(((((........))))).....)))))....)))...... (-22.66 = -22.90 + 0.24)



| Location | 23,906,881 – 23,906,975 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.35 |
| Mean single sequence MFE | -30.44 |
| Consensus MFE | -29.64 |
| Energy contribution | -30.32 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.97 |
| SVM decision value | 3.99 |
| SVM RNA-class probability | 0.999743 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23906881 94 - 27905053 AAUUUCCCAUUUUCAAUCCGUGGGCGUCGUGGGCCAGACACAAAUAAUUGAAGUCUGGUCAGCCCAAUGGCCAUUAUGGGACCA-AAUGGCCAGA ....................(((((......((((((((.(((....)))..)))))))).))))).((((((((.((....))-)))))))).. ( -34.90) >DroSec_CAF1 107442 89 - 1 -----CCCAUUUUCAAUCCGUGGGCGUCGUGGGCCAGACACAAAUAAUUGGAGUCUGGUCAGCCCAAUGGCCAUUAUGGGACCA-AAUGGCCAGA -----...............(((((......((((((((.(((....)))..)))))))).))))).((((((((.((....))-)))))))).. ( -36.40) >DroSim_CAF1 110135 91 - 1 ---UUCCCAUUUUCAAUCCGUGGGCGUCGUGGGCCAGACACAAAUAAUUGGAGUCUGGUCAGCCCAAUGGCUAUUAUGGGACCA-AAUGGCCAGA ---.................(((((......((((((((.(((....)))..)))))))).))))).((((((((.((....))-)))))))).. ( -34.10) >DroEre_CAF1 111496 94 - 1 AAUUUUCCAUUUUCAAUCCGUGGGCGUCGUGGGGCAGACACAAAUAAUUGGAGUCUGGUCAGUCCAAUGGCCAUUAUGGGACCA-AAUAGCCAGA ....((((((.......((((((((......((.(((((.(((....)))..))))).)).)))).)))).....))))))...-.......... ( -23.20) >DroYak_CAF1 115109 95 - 1 AAUUUCGCAUUUUCAAUCCGUGGGCGUCGUGGGGCAGACACAAAUAAUUGGAGUCUGGUCAGUGCAAUGGCCAUUAUGGGACCAAAAUAGCCAAA ......(((((..((...((....))...))((.(((((.(((....)))..))))).)))))))..((((.(((.((....)).))).)))).. ( -23.60) >consensus AAUUUCCCAUUUUCAAUCCGUGGGCGUCGUGGGCCAGACACAAAUAAUUGGAGUCUGGUCAGCCCAAUGGCCAUUAUGGGACCA_AAUGGCCAGA ....................(((((......((((((((.(((....)))..)))))))).))))).((((((((.((....)).)))))))).. (-29.64 = -30.32 + 0.68)

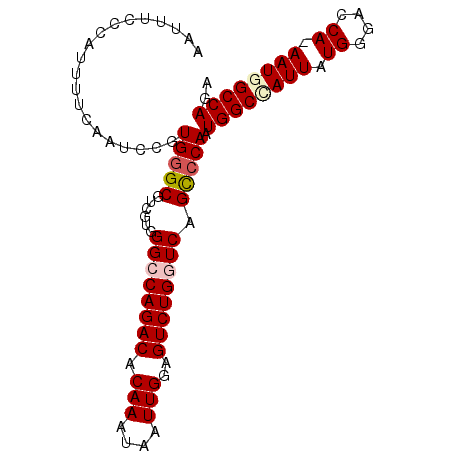

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:22:32 2006