
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,858,227 – 23,858,347 |
| Length | 120 |
| Max. P | 0.978694 |

| Location | 23,858,227 – 23,858,321 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 76.51 |
| Mean single sequence MFE | -32.14 |
| Consensus MFE | -15.44 |
| Energy contribution | -17.02 |
| Covariance contribution | 1.58 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.48 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.957543 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23858227 94 + 27905053 AGGACAAGCAGAUAGUGUCUGCUGUCCG----UUCGGCUCCUGAGUUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUC .(((((.((((((...))))))))))).----...(((.(..((((((..(((((....)))))..)))))).......((((.....)))).).))) ( -35.70) >DroSec_CAF1 63765 94 + 1 AGGACAAGCAGAUAGUGUCUGCUGUCCG----UUCGGCUCCUGAGCUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGUCGUGUC .(((((.((((((...))))))))))).----...(((....((((((..(((((....)))))..)))))).......)))................ ( -37.20) >DroSim_CAF1 63209 94 + 1 AGGACAAGCAGAUAUUGUCUGCUGUCCG----UUCGGCUCCUGAGCUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUC .(((((.((((((...))))))))))).----...(((.(..((((((..(((((....)))))..)))))).......((((.....)))).).))) ( -38.10) >DroEre_CAF1 67780 82 + 1 AGGACAAGG----AGUGUUUGCUGUCCG----UUCGGCUCCU--------GAGCUCCACAGCUCCACGGCUCCUUCCUUUCCAAUUCUUGGCCGUGUC ((((..(((----(((((..(((((..(----(((((...))--------))))...)))))...)).))))))))))..(((.....)))....... ( -24.20) >DroYak_CAF1 65727 90 + 1 AGGACAAGCAGAUAGUGUCUGCUGUCCGUCCAUUCGGCUACU--------GAGCUUCUCAGCUCCACAGCUCCUUCCUUUCCAAUGCUUGGCCGUGUC .(((((.((((((...))))))))))).......((((((..--------(((((....)))))...(((...............))))))))).... ( -29.66) >DroAna_CAF1 65429 79 + 1 AGGACAAGCAGAUAGCGGAUAGUGUCUG----UUCACUUAUCCUGCUGCUG-GCUUCCCGUCGGCCCUGCUC--------------CUUGGCCGUGUC .((.(((((((..(((((((((((....----..))).))))).)))((((-((.....)))))).))))).--------------..)).))..... ( -28.00) >consensus AGGACAAGCAGAUAGUGUCUGCUGUCCG____UUCGGCUCCUGAGCUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUC .(((((.(((((.....)))))))))).......................(((((....)))))(((.(((..................))).))).. (-15.44 = -17.02 + 1.58)

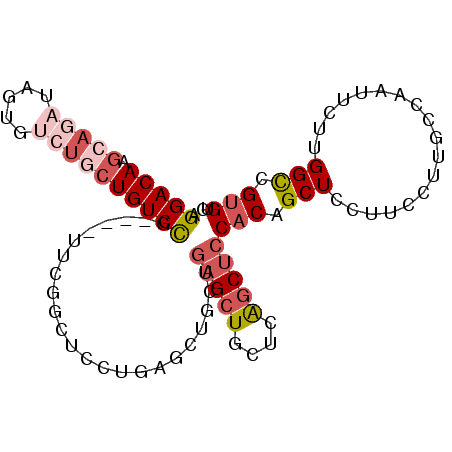

| Location | 23,858,227 – 23,858,321 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 76.51 |
| Mean single sequence MFE | -33.63 |
| Consensus MFE | -17.22 |
| Energy contribution | -19.58 |
| Covariance contribution | 2.36 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.51 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978694 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23858227 94 - 27905053 GACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAACUCAGGAGCCGAA----CGGACAGCAGACACUAUCUGCUUGUCCU ......((((.....))))..((...(((.(((.(((((....))))).))).))).....))...----.((((((((((.....))))).))))). ( -36.10) >DroSec_CAF1 63765 94 - 1 GACACGACCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAGCUCAGGAGCCGAA----CGGACAGCAGACACUAUCUGCUUGUCCU ..............(((((.......(((((((.(((((....))))).)))))))....))))).----.((((((((((.....))))).))))). ( -40.40) >DroSim_CAF1 63209 94 - 1 GACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAGCUCAGGAGCCGAA----CGGACAGCAGACAAUAUCUGCUUGUCCU ......((((.....))))..((...(((((((.(((((....))))).))))))).....))...----.((((((((((.....))))).))))). ( -42.70) >DroEre_CAF1 67780 82 - 1 GACACGGCCAAGAAUUGGAAAGGAAGGAGCCGUGGAGCUGUGGAGCUC--------AGGAGCCGAA----CGGACAGCAAACACU----CCUUGUCCU ..(((((((.....((....))....).))))))(((((....)))))--------..........----.((((((........----..)))))). ( -23.80) >DroYak_CAF1 65727 90 - 1 GACACGGCCAAGCAUUGGAAAGGAAGGAGCUGUGGAGCUGAGAAGCUC--------AGUAGCCGAAUGGACGGACAGCAGACACUAUCUGCUUGUCCU ..(((((((.....((....))....).))))))(((((....)))))--------....(((....)).)((((((((((.....))))).))))). ( -31.10) >DroAna_CAF1 65429 79 - 1 GACACGGCCAAG--------------GAGCAGGGCCGACGGGAAGC-CAGCAGCAGGAUAAGUGAA----CAGACACUAUCCGCUAUCUGCUUGUCCU (((((((((...--------------......)))))..((....)-)(((((..(((((.(((..----....)))))))).....))))))))).. ( -27.70) >consensus GACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAGCUCAGGAGCCGAA____CGGACAGCAGACACUAUCUGCUUGUCCU ..(((((((.....((....))....).))))))(((((....))))).......................((((((((((.....))))).))))). (-17.22 = -19.58 + 2.36)



| Location | 23,858,255 – 23,858,347 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 90.11 |
| Mean single sequence MFE | -23.77 |
| Consensus MFE | -16.66 |
| Energy contribution | -19.30 |
| Covariance contribution | 2.64 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585525 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23858255 92 + 27905053 --UUCGGCUCCUGAGUUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU --...((((...((((((..(((((....)))))..)))))).......((((.....)))).(((..(((....)))..))).....)))).. ( -26.20) >DroSec_CAF1 63793 92 + 1 --UUCGGCUCCUGAGCUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGUCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU --...((((...((((((..(((((....)))))..)))))).......((.((((.((((......)))).)))).)).........)))).. ( -27.70) >DroSim_CAF1 63237 92 + 1 --UUCGGCUCCUGAGCUGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU --...((((...((((((..(((((....)))))..)))))).......((((.....)))).(((..(((....)))..))).....)))).. ( -28.60) >DroEre_CAF1 67804 84 + 1 --UUCGGCUCCU--------GAGCUCCACAGCUCCACGGCUCCUUCCUUUCCAAUUCUUGGCCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU --...((((...--------(((((....)))))(((((((..................)))))))......................)))).. ( -20.07) >DroYak_CAF1 65757 86 + 1 CAUUCGGCUACU--------GAGCUUCUCAGCUCCACAGCUCCUUCCUUUCCAAUGCUUGGCCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU .....(((((((--------(((...)))))..(((.(((...............))))))..(((..(((....)))..)))....))))).. ( -16.26) >consensus __UUCGGCUCCUGAG_UGCUGAGCUGCUCAGCUCCACAGCUCCUUCCUUGCCAAUUCUUGGCCGUGUCCGACGAUUCGUUUAUUUGAUAGCCUU .....((((...((((((..(((((....)))))..)))))).......((((.....)))).(((..(((....)))..))).....)))).. (-16.66 = -19.30 + 2.64)

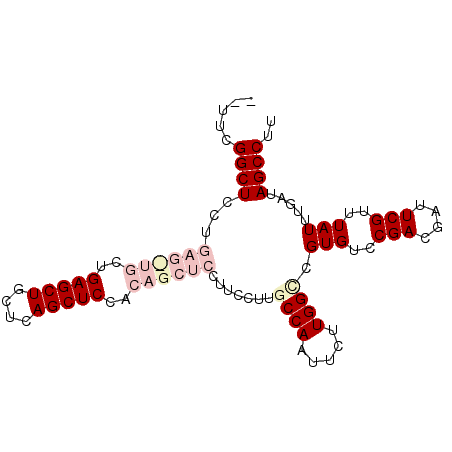

| Location | 23,858,255 – 23,858,347 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 90.11 |
| Mean single sequence MFE | -29.44 |
| Consensus MFE | -21.52 |
| Energy contribution | -21.56 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.12 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965502 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23858255 92 - 27905053 AAGGCUAUCAAAUAAACGAAUCGUCGGACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAACUCAGGAGCCGAA-- ..((((.((.......(((....)))......((((.....))))...))..(((.(((.(((((....))))).))).)))...))))...-- ( -27.50) >DroSec_CAF1 63793 92 - 1 AAGGCUAUCAAAUAAACGAAUCGUCGGACACGACCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAGCUCAGGAGCCGAA-- ..((((.((.......(((.((((((....))))...)).))).........(((((((.(((((....))))).))))))).))))))...-- ( -33.60) >DroSim_CAF1 63237 92 - 1 AAGGCUAUCAAAUAAACGAAUCGUCGGACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCAGCUCAGGAGCCGAA-- ..((((.((.......(((....)))......((((.....))))...))..(((((((.(((((....))))).)))))))...))))...-- ( -34.10) >DroEre_CAF1 67804 84 - 1 AAGGCUAUCAAAUAAACGAAUCGUCGGACACGGCCAAGAAUUGGAAAGGAAGGAGCCGUGGAGCUGUGGAGCUC--------AGGAGCCGAA-- ..((((.((.......(((....)))..(((((((.....((....))....).))))))(((((....)))))--------.))))))...-- ( -25.90) >DroYak_CAF1 65757 86 - 1 AAGGCUAUCAAAUAAACGAAUCGUCGGACACGGCCAAGCAUUGGAAAGGAAGGAGCUGUGGAGCUGAGAAGCUC--------AGUAGCCGAAUG ..((((((........(((....)))..(((((((.....((....))....).))))))(((((....)))))--------.))))))..... ( -26.10) >consensus AAGGCUAUCAAAUAAACGAAUCGUCGGACACGGCCAAGAAUUGGCAAGGAAGGAGCUGUGGAGCUGAGCAGCUCAGCA_CUCAGGAGCCGAA__ ..((((..........(((....)))..(((((((.....((....))....).))))))(((((....)))))...........))))..... (-21.52 = -21.56 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:22:14 2006