
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,749,690 – 23,749,793 |
| Length | 103 |
| Max. P | 0.733872 |

| Location | 23,749,690 – 23,749,793 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 74.70 |
| Mean single sequence MFE | -29.22 |
| Consensus MFE | -18.01 |
| Energy contribution | -17.93 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.650730 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
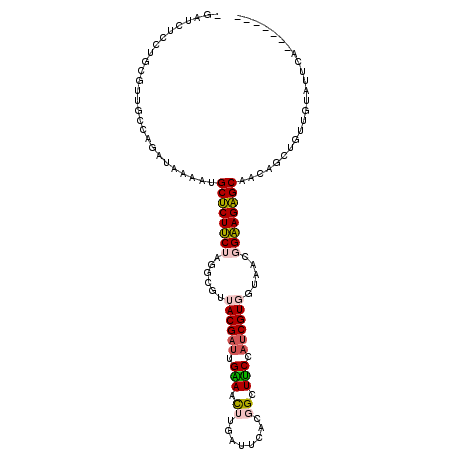
>3R_DroMel_CAF1 23749690 103 + 27905053 -GAUCUCCUGCGUUGCCAGAUAAAAUGCUCUUCUUGGCGUUACGAUUGAAACUUGAUUCACGGCUUCCAUCGUGGUAACGGAAGAGCAACAGCUGUUGUAUUCA------- -.........(((((((......((((((......))))))(((((.(((.((........)).))).))))))))))))(((..(((((....))))).))).------- ( -27.80) >DroSec_CAF1 2698 103 + 1 -GAUCUCCUGCGUUGUCAGAUAAAAUGCUCUUCUUGGCGUUACGAUUGAAAUUUGAUUCACGGCUUCCAUCGUGGUAACGGAAGAGCAACAGCUGUAGUAUUCA------- -((....(((((((((.........((((((((....(((((((((..(...)..)))(((((......)))))))))))))))))))))))).))))...)).------- ( -28.40) >DroSim_CAF1 2617 103 + 1 -GAUCUCCUGCGUUGCCAGAUAAAAUGCUCUUCUUGGCGUUACGAUUGAAAUUUGAUUCACGGCUUCCAUCGUGGUAACGGAAGAGCAACAGCUGUUGUAUUUA------- -...(((.(.(((((((......((((((......))))))(((((.(((..(((.....))).))).)))))))))))).).)))((((....))))......------- ( -25.20) >DroYak_CAF1 868 110 + 1 -GAUAUCCUGCGUUGCCAGAUAAAAUGCUCUUCUAGGCGUUACGAUUGAAACUUGUUUCACGGCUUCCAUCGUGGUAACGGAAGAGCUACAUCUUUUGAACCCUUUCAAAC -....((.(.(((((((......((((((......))))))(((((.(((.((........)).))).)))))))))))).).)).........((((((....)))))). ( -25.00) >DroMoj_CAF1 2129 98 + 1 AGCAAUAAUGCGUUGCUAAAUAAACUGCCCUCCUAGGCGUUACGAUUGCAGCCUGAUUCCAAGGUGCCAUCGUUUGCAGUGGAGGGCCACAGCUGCA---U---------- ((((((.....)))))).........(((((((...(((..(((((.(((.(((.......)))))).))))).)))...)))))))..........---.---------- ( -37.10) >DroPer_CAF1 2589 102 + 1 AGCAUUGGUGCGUUGCCAGAAUAACUGCUCUCCUAGGCGUUACGAGUGUAACCUGAUUCAAAGUUGCAGACGUUUGCAGUGGAGAGCCACAGCUU--AUUUUUA------- (((.(((((.....))))).......(((((((...(((..(((..((((((.(......).))))))..))).)))...)))))))....))).--.......------- ( -31.80) >consensus _GAUCUCCUGCGUUGCCAGAUAAAAUGCUCUUCUAGGCGUUACGAUUGAAACUUGAUUCACGGCUUCCAUCGUGGUAACGGAAGAGCAACAGCUGUUGUAUUCA_______ ..........................((((((((......((((((.(((.((........)).))).)))))).....))))))))........................ (-18.01 = -17.93 + -0.08)



| Location | 23,749,690 – 23,749,793 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 74.70 |
| Mean single sequence MFE | -27.10 |
| Consensus MFE | -17.96 |
| Energy contribution | -18.24 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.733872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
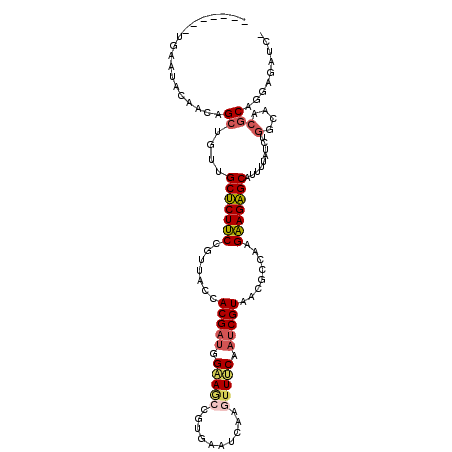
>3R_DroMel_CAF1 23749690 103 - 27905053 -------UGAAUACAACAGCUGUUGCUCUUCCGUUACCACGAUGGAAGCCGUGAAUCAAGUUUCAAUCGUAACGCCAAGAAGAGCAUUUUAUCUGGCAACGCAGGAGAUC- -------............((((((((((((((((...(((((.(((((..........))))).)))))))))....))))))))........(....)))))......- ( -25.80) >DroSec_CAF1 2698 103 - 1 -------UGAAUACUACAGCUGUUGCUCUUCCGUUACCACGAUGGAAGCCGUGAAUCAAAUUUCAAUCGUAACGCCAAGAAGAGCAUUUUAUCUGACAACGCAGGAGAUC- -------.........(((.((.((((((((((((((....((((...))))(((......)))....))))))....))))))))...)).)))...............- ( -22.00) >DroSim_CAF1 2617 103 - 1 -------UAAAUACAACAGCUGUUGCUCUUCCGUUACCACGAUGGAAGCCGUGAAUCAAAUUUCAAUCGUAACGCCAAGAAGAGCAUUUUAUCUGGCAACGCAGGAGAUC- -------......((((....))))(((((.((((.(((.(((((((((..((((......))))...))...((........)).)))))))))).)))).)))))...- ( -24.00) >DroYak_CAF1 868 110 - 1 GUUUGAAAGGGUUCAAAAGAUGUAGCUCUUCCGUUACCACGAUGGAAGCCGUGAAACAAGUUUCAAUCGUAACGCCUAGAAGAGCAUUUUAUCUGGCAACGCAGGAUAUC- .((((((....)))))).(((((.(((((((((((...(((((.(((((..........))))).)))))))))....)))))))......((((......)))))))))- ( -27.60) >DroMoj_CAF1 2129 98 - 1 ----------A---UGCAGCUGUGGCCCUCCACUGCAAACGAUGGCACCUUGGAAUCAGGCUGCAAUCGUAACGCCUAGGAGGGCAGUUUAUUUAGCAACGCAUUAUUGCU ----------.---.((((.(((((((((((...((..(((((.(((.((((....)))).))).)))))...))...))))))).(((.....)))..))))...)))). ( -36.30) >DroPer_CAF1 2589 102 - 1 -------UAAAAAU--AAGCUGUGGCUCUCCACUGCAAACGUCUGCAACUUUGAAUCAGGUUACACUCGUAACGCCUAGGAGAGCAGUUAUUCUGGCAACGCACCAAUGCU -------.......--.(((.((((((((((..((((......))))............(((((....))))).....))))))).........(....).)))....))) ( -26.90) >consensus _______UGAAUACAACAGCUGUUGCUCUUCCGUUACCACGAUGGAAGCCGUGAAUCAAGUUUCAAUCGUAACGCCAAGAAGAGCAUUUUAUCUGGCAACGCAGGAGAUC_ ..................((....(((((((.......(((((.(((((..........))))).)))))........))))))).........(....)))......... (-17.96 = -18.24 + 0.28)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:21:21 2006