| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
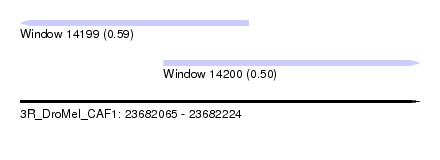
| Location | 23,682,065 – 23,682,224 |
| Length | 159 |
| Max. P | 0.585079 |

| Location | 23,682,065 – 23,682,156 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 94.95 |
| Mean single sequence MFE | -22.48 |
| Consensus MFE | -17.14 |
| Energy contribution | -18.10 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585079 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23682065 91 - 27905053 UUCCUUUUGUCGCUCUGUCACUUGAAUAAAAGGCGUAAAUCCGUUUUUUAUCGUCCUCUGGUUCCCGUGAUUUCCAUGGCCUUGAAAGGGA (((((((((..(((..(((((..((((...(((((((((.......)))).)).)))...))))..)))))......)))..))))))))) ( -23.10) >DroSec_CAF1 143600 91 - 1 UUCCUUUUAUCGCUCUGUCACUUGAAUCAAAGGCGUAAAUCCGUUUUUUAUCGUCCUCUGGCUUCUGUGAUUUCCAUGGCCUUGAAAGGGA (((((((((..(((..(((((..((((((.(((((((((.......)))).)).))).))).))).)))))......)))..))))))))) ( -25.00) >DroSim_CAF1 143471 90 - 1 UUCCUUUUGUCGCUCUGUCACUUGAAUCAAAGGCGUAAAUCCGUUUUUUAUCGUC-UCUGGCUCCUGUGAUUUCCAUGGCCUUGAAAGCGA .((((((((..(((..(((((..((..((.(((((((((.......)))).))))-).))..))..)))))......)))..)))))).)) ( -22.30) >DroEre_CAF1 150588 91 - 1 UUCCUUUUGUCGCUCUGUCACUUAAAUCAAAGGCGUAAAUCCGUUUUUUAUCGUCCUCUGGCUCCUGUGAUUUCCAUGCCCUUGAAAGGGA (((((((((..((...(((((.....(((.(((((((((.......)))).)).))).))).....)))))......))...))))))))) ( -20.00) >DroYak_CAF1 150191 91 - 1 UUCCUUUUGUCGCUCUGUCACUUGAAUCAAAGGCGUAAAUCCGUUUUUUAUCGUCCUCCGGCUCCUGUGAUUUCCAUGCCCUUGAAAGGAA .((((((((..((...(((((..((..(..(((((((((.......)))).)).)))..)..))..)))))......))...)))))))). ( -22.00) >consensus UUCCUUUUGUCGCUCUGUCACUUGAAUCAAAGGCGUAAAUCCGUUUUUUAUCGUCCUCUGGCUCCUGUGAUUUCCAUGGCCUUGAAAGGGA (((((((((..(((..(((((..((..((.(((((((((.......)))).)).))).))..))..)))))......)))..))))))))) (-17.14 = -18.10 + 0.96)



| Location | 23,682,122 – 23,682,224 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.56 |
| Mean single sequence MFE | -20.97 |
| Consensus MFE | -14.14 |
| Energy contribution | -14.19 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
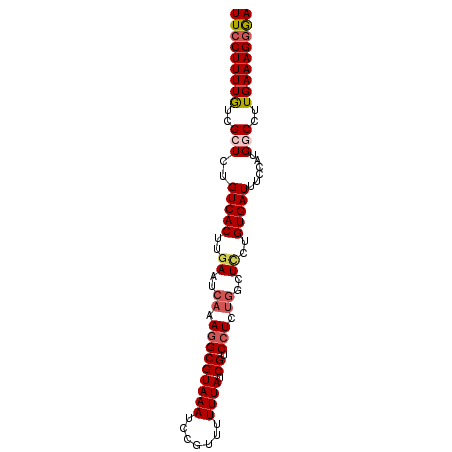
>3R_DroMel_CAF1 23682122 102 + 27905053 GCCUUUUAUU-CAAGUGACAGAGCGACA---AAAGGAA----------CAGGACU--CCGUUGAACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG--UACAAUAAAUGGCAAAAUAA (((.((((((-((.(((.(((.(((((.---...(((.----------......)--))(((((.((.....)))))))....)))))))))))..))--...)))))).)))....... ( -26.90) >DroPse_CAF1 152899 112 + 1 GCCUUUGAUU-UAAGUGACAGGGCGACA---AAAGAAACAAAAAGAAACAAGGCA--GCGGGACACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG--UACAAUAAAUGGCAAAAUAA (((.(((((.-...(((...(((((((.---.......(.....)......((((--.(........)....)))).......))))))).)))...)--).))).....)))....... ( -20.40) >DroSec_CAF1 143657 102 + 1 GCCUUUGAUU-CAAGUGACAGAGCGAUA---AAAGGAA----------CCGGACU--CCGUUGAACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG--UACAAUAAAUGGCAAAAUAA (((.(((((.-...(((.(((.(((((.---...(((.----------......)--))(((((.((.....)))))))....)))))))))))...)--).))).....)))....... ( -22.50) >DroWil_CAF1 159528 112 + 1 GCCUUUGCUU-UAAGUGACAGCGCGACGGGGAAAUGUAUAAAAAGAA---AGUGA--ACAGAAAACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG--UACAAUAAAUGGCAAAAUAA (((.(((...-...(((.(((.(((((....................---.(...--.).....(((.....)))........)))))))))))....--..))).....)))....... ( -18.60) >DroAna_CAF1 149324 107 + 1 GCUUUUGAUUUUAAGUGACAGAGCGACA---AAAGGAA----------CAGGACUGGCGAACGAAAAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUGUGUACAAUAAAUGGCAAAAUAA (((.(((((.....(((.(((.(((((.---..((...----------.....))((((.............)))).......))))))))))).....)).))).....)))....... ( -17.02) >DroPer_CAF1 149124 112 + 1 GCCUUUGAUU-UAAGUGACAGGGCGACA---AAAGAAACAAAAAGAAACAAGGCA--GCGGGACACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG--UACAAUAAAUGGCAAAAUAA (((.(((((.-...(((...(((((((.---.......(.....)......((((--.(........)....)))).......))))))).)))...)--).))).....)))....... ( -20.40) >consensus GCCUUUGAUU_UAAGUGACAGAGCGACA___AAAGGAA__________CAAGACA__CCGGGAAACAGAUAAUGUCAAUAUUAGUCGCCUGCAUAAUG__UACAAUAAAUGGCAAAAUAA (((.(((.......(((.(((.(((((........................(......)..........(((((....))))))))))))))))........))).....)))....... (-14.14 = -14.19 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:21:05 2006