
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,681,113 – 23,681,228 |
| Length | 115 |
| Max. P | 0.997364 |

| Location | 23,681,113 – 23,681,206 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 85.33 |
| Mean single sequence MFE | -32.20 |
| Consensus MFE | -20.91 |
| Energy contribution | -21.97 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -4.05 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.977930 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23681113 93 + 27905053 UCCAGGACAUCAGGAUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUGCUGUUCCUGCUGCUGCUGUGU ..(((.(((.((((((((((((((.......))))....)))))....((((((((((((....))))))))))))))))).)).).)))... ( -32.80) >DroSec_CAF1 142724 87 + 1 UCCAUGACAUCAGGAUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUG------CUGUUGCUGCUGUGU ((((.((.((....)).))))))............((((((....(((((((((((((((....))))))))------)))))))..)))))) ( -35.70) >DroSim_CAF1 142575 93 + 1 UCCAGGACAUCAGGAUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUGCUGUUGCUGUUGCUGCUGUGU ((((((...........))))))............((((((....((((((((((((((((.......).)))))))))))))))..)))))) ( -37.00) >DroYak_CAF1 149261 72 + 1 UUCAGGAUAUCAGGAUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCU---------------------GCUGCUGUGU (((.((((((....)))))).)))..........(((........)))(((((((((....)---------------------)))))))).. ( -23.30) >consensus UCCAGGACAUCAGGAUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUG______CUGUUGCUGCUGUGU ((((.((.((....)).))))))...........(((........)))((((((((((((.....((....))......)))))))))))).. (-20.91 = -21.97 + 1.06)

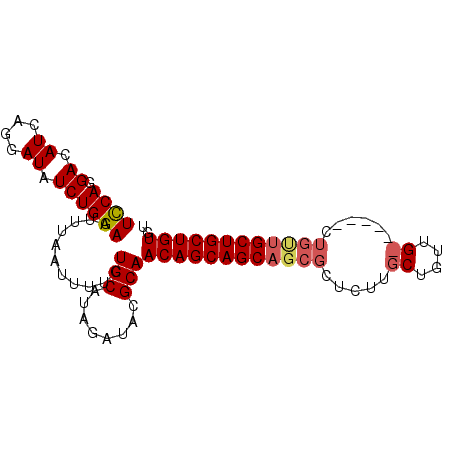

| Location | 23,681,113 – 23,681,206 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 85.33 |
| Mean single sequence MFE | -29.71 |
| Consensus MFE | -21.20 |
| Energy contribution | -21.51 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -4.12 |
| Structure conservation index | 0.71 |
| SVM decision value | 2.30 |
| SVM RNA-class probability | 0.991926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23681113 93 - 27905053 ACACAGCAGCAGCAGGAACAGCAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAUCCUGAUGUCCUGGA ..(((((((((((.(......)....((....)))))))))))))(((((....)))))...........(((((((((....)))).))))) ( -32.40) >DroSec_CAF1 142724 87 - 1 ACACAGCAGCAACAG------CAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAUCCUGAUGUCAUGGA .....((((((((((------((.((((......)))).))))))))).......)))............(((((((((....))))).)))) ( -30.51) >DroSim_CAF1 142575 93 - 1 ACACAGCAGCAACAGCAACAGCAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAUCCUGAUGUCCUGGA .....(((......(((((((((.((((......)))).))))))))).......)))............(((((((((....)))).))))) ( -33.82) >DroYak_CAF1 149261 72 - 1 ACACAGCAGC---------------------AGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAUCCUGAUAUCCUGAA ..((((((((---------------------(....)))))))))(((((....)))))..........(((..(((((....)))))..))) ( -22.10) >consensus ACACAGCAGCAACAG______CAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAUCCUGAUGUCCUGGA ..(((((((((.........................)))))))))(((((....)))))...........(((((((((....))))).)))) (-21.20 = -21.51 + 0.31)



| Location | 23,681,127 – 23,681,228 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 85.57 |
| Mean single sequence MFE | -31.06 |
| Consensus MFE | -20.95 |
| Energy contribution | -23.07 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.67 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.982019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23681127 101 + 27905053 AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUGCUGUUCCUGCUGCUGCUGUGUGCGAUAUGAAAAAUGCAAAGGG ...(((((((......))))(((((..(((((((((((((((((((.....((....))......)))))))))))).)))).))).....))))).))). ( -33.60) >DroSec_CAF1 142738 95 + 1 AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUG------CUGUUGCUGCUGUGUGCGAUAUGAAAAAUGCAAAGGG (((((.((((.......))))((((((....(((((((((((((((....))))))))------)))))))..))))))..)))))............... ( -36.60) >DroSim_CAF1 142589 101 + 1 AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUGCUGUUGCUGUUGCUGCUGUGUGCGAUAUGAAAAAUGCAAAGGG ...(((((((......))))(((((......(((((((((((((((....)))))))))))))))((((((........))))))......))))).))). ( -37.30) >DroEre_CAF1 149669 77 + 1 AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCU---------------------G---CUGUGUGCGAUAUGAAAAAUGCAAAGGG ...(((((((......))))(((((..(((((((((((((((...)))---------------------)---)))).)))).))).....))))).))). ( -22.20) >DroYak_CAF1 149275 80 + 1 AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCU---------------------GCUGCUGUGUGCGAUAUGAAAAAUGCAAAGGG ...(((((((......))))(((((..((((((((((((((((....)---------------------)))))))).)))).))).....))))).))). ( -25.60) >consensus AUAUCUGGAAUUUAAUUUUCUGCAUAGAUACGCAACAGCAGCAGCGCUCUUGCUGUUG______CUG_UGCUGCUGUGUGCGAUAUGAAAAAUGCAAAGGG ...(((((((......))))(((((..(((((((((((((((((((...................)))))))))))).)))).))).....))))).))). (-20.95 = -23.07 + 2.12)



| Location | 23,681,127 – 23,681,228 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 85.57 |
| Mean single sequence MFE | -29.28 |
| Consensus MFE | -20.81 |
| Energy contribution | -21.81 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.53 |
| Structure conservation index | 0.71 |
| SVM decision value | 2.85 |
| SVM RNA-class probability | 0.997364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23681127 101 - 27905053 CCCUUUGCAUUUUUCAUAUCGCACACAGCAGCAGCAGGAACAGCAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((.....(((.((((.(((((((((((.(......)....((....))))))))))))))))))))..))))))).................. ( -33.90) >DroSec_CAF1 142738 95 - 1 CCCUUUGCAUUUUUCAUAUCGCACACAGCAGCAACAG------CAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((....(.....((.....)).(((((((------((.((((......)))).)))))))))).....))))))).................. ( -30.30) >DroSim_CAF1 142589 101 - 1 CCCUUUGCAUUUUUCAUAUCGCACACAGCAGCAACAGCAACAGCAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((.....(((.((((.(((((((((.(.((....))....((....))).))))))))))))))))..))))))).................. ( -30.60) >DroEre_CAF1 149669 77 - 1 CCCUUUGCAUUUUUCAUAUCGCACACAG---C---------------------AGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((.....(((.((((.((((---(---------------------(((...)))))))))))))))..))))))).................. ( -24.10) >DroYak_CAF1 149275 80 - 1 CCCUUUGCAUUUUUCAUAUCGCACACAGCAGC---------------------AGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((.....(((.((((.((((((((---------------------(....))))))))))))))))..))))))).................. ( -27.50) >consensus CCCUUUGCAUUUUUCAUAUCGCACACAGCAGCA_CAG______CAACAGCAAGAGCGCUGCUGCUGUUGCGUAUCUAUGCAGAAAAUUAAAUUCCAGAUAU ...(((((((.....(((.((((.(((((((((.........................))))))))))))))))..))))))).................. (-20.81 = -21.81 + 1.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:21:03 2006