
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,647,478 – 23,647,571 |
| Length | 93 |
| Max. P | 0.890051 |

| Location | 23,647,478 – 23,647,571 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 77.67 |
| Mean single sequence MFE | -22.28 |
| Consensus MFE | -9.23 |
| Energy contribution | -9.40 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.41 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.890051 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23647478 93 + 27905053 AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCA--CAAAACUGCAA------CACA-UUGCAGCUGGAACAGCACCAGAGGCAAACUAACU ........((((((........)))))).....((((((.(((.--.....((((((------....-))))))((((.......)))).)))...)))))) ( -24.50) >DroVir_CAF1 148860 84 + 1 AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCA--CAAAACUGUAACUGCAGCACA-CUGCUG---CAUCACAGGCC---------CA--- .((...(((((((((......((((.....))))))))))).))--.....((((...((((((...-..))))---))..))))..)---------).--- ( -24.70) >DroPse_CAF1 111993 82 + 1 AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCACACAAAAGUGUAA------CACAGCUGCAGCUGUAACAGAA--------------ACA .(....(((((((((......((((.....))))))))))((.((((....)))).)------)(((((....))))).)))...--------------.). ( -20.30) >DroMoj_CAF1 158276 84 + 1 AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCA--CAAAACUGUAACUGCAGCACG-CUGCGG---CAUCACAGGCU---------CG--- ......(((((((((......((((.....))))))))))).))--.....((((..((((((....-))))))---....))))...---------..--- ( -21.30) >DroAna_CAF1 110420 84 + 1 GGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCA--CAAAACUGCAA------CACA-CUGCAGCUGGAACAGAAGCG---------CCACU ........((((((........)))))).....(((.(((((..--.....(((((.------....-.)))))(((...)))..)))---------))))) ( -22.60) >DroPer_CAF1 110321 82 + 1 AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCACACAAAAGUGUAA------CACAGCUGCAGCUGUAACAGAA--------------ACA .(....(((((((((......((((.....))))))))))((.((((....)))).)------)(((((....))))).)))...--------------.). ( -20.30) >consensus AGGAUAUGUGCCAAUAAAAAGUUUGGCAAACAAAGUUGGCGUCA__CAAAACUGUAA______CACA_CUGCAGCUGCAACAGAAGC__________C_AC_ ......(((((((((......((((.....))))))))))).)).......(((((.............)))))............................ ( -9.23 = -9.40 + 0.17)


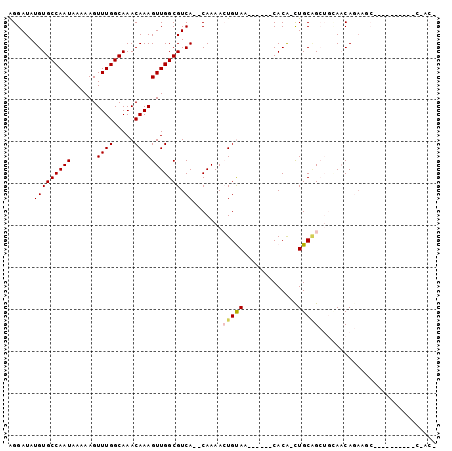
| Location | 23,647,478 – 23,647,571 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 77.67 |
| Mean single sequence MFE | -23.83 |
| Consensus MFE | -7.50 |
| Energy contribution | -7.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.31 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513479 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23647478 93 - 27905053 AGUUAGUUUGCCUCUGGUGCUGUUCCAGCUGCAA-UGUG------UUGCAGUUUUG--UGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ((((...((((..((((.......))))..))))-.(((------(..(......)--..)))).)))).(((.((((((........)))))).))).... ( -24.00) >DroVir_CAF1 148860 84 - 1 ---UG---------GGCCUGUGAUG---CAGCAG-UGUGCUGCAGUUACAGUUUUG--UGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ---..---------(((......((---((((..-...))))))(((((......)--))))))).....(((.((((((........)))))).))).... ( -27.30) >DroPse_CAF1 111993 82 - 1 UGU--------------UUCUGUUACAGCUGCAGCUGUG------UUACACUUUUGUGUGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ...--------------....((((((((....)))))(------((((((....)))))))...)))..(((.((((((........)))))).))).... ( -23.10) >DroMoj_CAF1 158276 84 - 1 ---CG---------AGCCUGUGAUG---CCGCAG-CGUGCUGCAGUUACAGUUUUG--UGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ---((---------((.(((((((.---..((((-....)))).))))))).))))--............(((.((((((........)))))).))).... ( -22.50) >DroAna_CAF1 110420 84 - 1 AGUGG---------CGCUUCUGUUCCAGCUGCAG-UGUG------UUGCAGUUUUG--UGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCC ..(((---------((..((......((((((((-....------))))))))...--.)))))))....(((.((((((........)))))).))).... ( -23.00) >DroPer_CAF1 110321 82 - 1 UGU--------------UUCUGUUACAGCUGCAGCUGUG------UUACACUUUUGUGUGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ...--------------....((((((((....)))))(------((((((....)))))))...)))..(((.((((((........)))))).))).... ( -23.10) >consensus _GU_G__________GCUUCUGUUACAGCUGCAG_UGUG______UUACAGUUUUG__UGACGCCAACUUUGUUUGCCAAACUUUUUAUUGGCACAUAUCCU ......................................................................(((.((((((........)))))).))).... ( -7.50 = -7.50 + 0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:20:50 2006