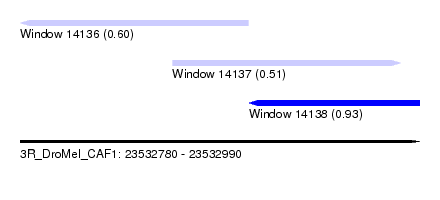
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,532,780 – 23,532,990 |
| Length | 210 |
| Max. P | 0.932257 |

| Location | 23,532,780 – 23,532,900 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -33.94 |
| Consensus MFE | -30.24 |
| Energy contribution | -31.13 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.595207 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
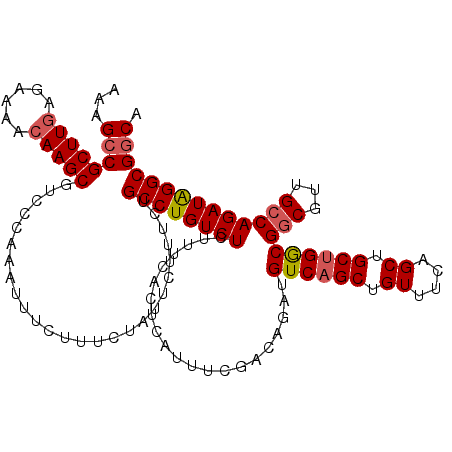
>3R_DroMel_CAF1 23532780 120 - 27905053 AAAGCCGCUUGAGAAAACAAGCGUCCCAAAUUUCUUUCUAUUCUUUCUGCCUGUCUCUUUCACUCAUUUCGACAGAUGUCAGCUGUUUCAGCUGCUGGCGGCGUUGCCAGAUAGGCGUCA .....((((((......)))))).........................((((((((.....................((((((.((....)).))))))(((...))))))))))).... ( -33.60) >DroSim_CAF1 567 120 - 1 AAAGCCGCUUGAGAAAACAAGCGUCCCAAAUUUCUUUCUAUUCUUUCUGCCUGUCUCUUUCACUAAUUUCAACAGAUGUCAGCUGUUUCAGCUGCCGACGACGCUGCCAGAUAGGCGGCA ...((((((((........((((((....................((((..((................)).))))((.((((((...)))))).))..))))))......)))))))). ( -31.63) >DroYak_CAF1 605 120 - 1 AAAGCCGCUUAAGAAAGCAAGCGUCCCAAAUUUCUUUCUUUUCUUUCUGCCUGUCUCUUUCACUCAUUUCGUCAGAUGUCAGCUGUUUCAGCUGCUGGCGGCGUUGCCAGAUGGGCGGCA ...(((....(((((((.((..........)).)))))))........((((((((..............(((((....((((((...)))))))))))(((...)))))))))))))). ( -36.60) >consensus AAAGCCGCUUGAGAAAACAAGCGUCCCAAAUUUCUUUCUAUUCUUUCUGCCUGUCUCUUUCACUCAUUUCGACAGAUGUCAGCUGUUUCAGCUGCUGGCGGCGUUGCCAGAUAGGCGGCA ...((((((((......)))))..........................((((((((.....................((((((.((....)).))))))(((...)))))))))))))). (-30.24 = -31.13 + 0.89)


| Location | 23,532,860 – 23,532,980 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.61 |
| Mean single sequence MFE | -33.53 |
| Consensus MFE | -30.95 |
| Energy contribution | -31.40 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.92 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.507320 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
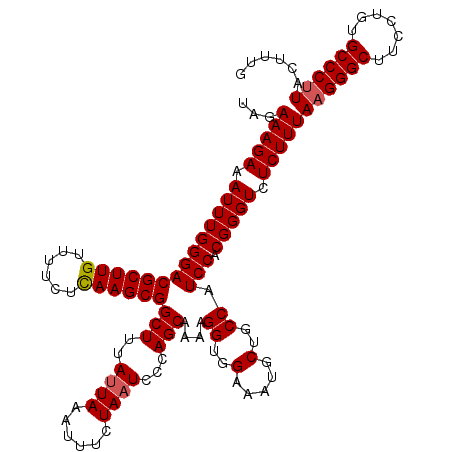
>3R_DroMel_CAF1 23532860 120 + 27905053 UAGAAAGAAAUUUGGGACGCUUGUUUUCUCAAGCGGCUUUAUUAAAUUUCUAAUCCCAGCAAGAGGUGGAAAUGCUGCCAUCCACGGGUCUCUUUAAGGGCUUCUUGUGCCCUUACUUUG ...(((((.((((((((.(((((......)))))(((..((((..((((((..........))))))...))))..))).))).))))).)))))((((((.......))))))...... ( -35.20) >DroSim_CAF1 647 120 + 1 UAGAAAGAAAUUUGGGACGCUUGUUUUCUCAAGCGGCUUUAUUAAAUUUCUAAUCCCAGCAAAAGGUGGAAAUGCUGCCAUCCACGGGUCUCUUUAAGGGCUUCGUGUGCCCUUACUUUG ...(((((.((((((((((((((......))))))(((..((((......))))...)))....((..(.....)..)).))).))))).)))))((((((.......))))))...... ( -35.00) >DroYak_CAF1 685 120 + 1 AAGAAAGAAAUUUGGGACGCUUGCUUUCUUAAGCGGCUUUAAUAAACAUCUAAUCCCAGCAAAAGGUGGAAAUGCUUCCAUCCACGGGUCUCUUUAAGGGCUUUCUGUGCCCGUACUUUG ...((((......((((((((.((((....))))((..(((.........)))..)))))....(((((((....))))))).....))))).....((((.......))))...)))). ( -30.40) >consensus UAGAAAGAAAUUUGGGACGCUUGUUUUCUCAAGCGGCUUUAUUAAAUUUCUAAUCCCAGCAAAAGGUGGAAAUGCUGCCAUCCACGGGUCUCUUUAAGGGCUUCCUGUGCCCUUACUUUG ...(((((.((((((((((((((......))))))(((..((((......))))...)))....((..(.....)..)).))).))))).)))))((((((.......))))))...... (-30.95 = -31.40 + 0.45)

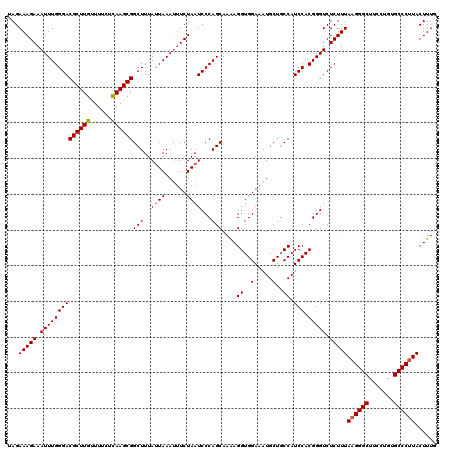
| Location | 23,532,900 – 23,532,990 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 93.70 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -27.67 |
| Energy contribution | -28.00 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932257 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23532900 90 - 27905053 UCCUGGCCGUCAAAGUAAGGGCACAAGAAGCCCUUAAAGAGACCCGUGGAUGGCAGCAUUUCCACCUCUUGCUGGGAUUAGAAAUUUAAU (((..((........(((((((.......)))))))(((((....(((((..........))))))))))))..)))............. ( -29.00) >DroSim_CAF1 687 90 - 1 UCCUGGCCGUCAAAGUAAGGGCACACGAAGCCCUUAAAGAGACCCGUGGAUGGCAGCAUUUCCACCUUUUGCUGGGAUUAGAAAUUUAAU (((..((.(((....(((((((.......)))))))....)))..(((((..........))))).....))..)))............. ( -29.00) >DroYak_CAF1 725 90 - 1 UCCUGGCCGUCAAAGUACGGGCACAGAAAGCCCUUAAAGAGACCCGUGGAUGGAAGCAUUUCCACCUUUUGCUGGGAUUAGAUGUUUAUU (((..((.(((....((.((((.......)))).))....)))..(((((((....))..))))).....))..)))............. ( -25.90) >consensus UCCUGGCCGUCAAAGUAAGGGCACAAGAAGCCCUUAAAGAGACCCGUGGAUGGCAGCAUUUCCACCUUUUGCUGGGAUUAGAAAUUUAAU (((..((.(((....(((((((.......)))))))....)))..(((((..........))))).....))..)))............. (-27.67 = -28.00 + 0.33)

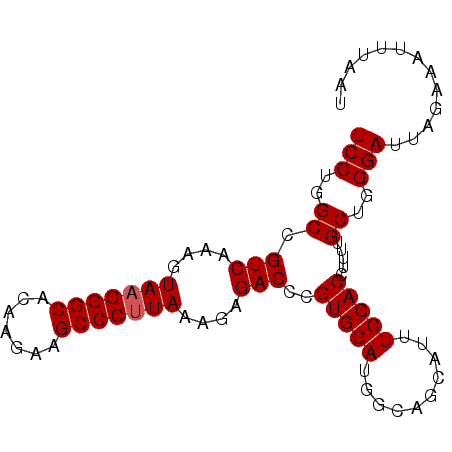
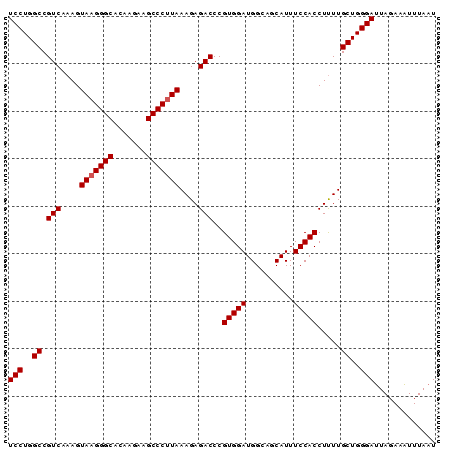
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:20:07 2006