
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,408,841 – 23,409,038 |
| Length | 197 |
| Max. P | 0.784992 |

| Location | 23,408,841 – 23,408,958 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 77.89 |
| Mean single sequence MFE | -42.06 |
| Consensus MFE | -23.23 |
| Energy contribution | -26.20 |
| Covariance contribution | 2.97 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784992 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23408841 117 - 27905053 CCCUCGAGGGCCAGUUCCAGGCACAUACUGGAUCCUCCGUCGGAUCCGGAACCCUGCUGGGAGUUGCCGUCGUGGGAAGUAGUUCCUCCUCCGCCCGAUCCCAGUCCCAAACUGGAC (((.(((.(((.(.((((((.((....((((((((......)))))))).....)))))))).).))).))).)))........................(((((.....))))).. ( -45.90) >DroGri_CAF1 46105 93 - 1 CCCUCCAACGCCAGUUCCAGGCACAUGCUUGAGCCACCAUCGGAU------CCAUGAUGCGAAUUG---------GAAUCGGGUCCACU---GCC------AAUUCCCAUUGUGGAC ...(((.......((((.((((....)))))))).......)))(------(((..(((.((((((---------(...(((.....))---)))------))))).)))..)))). ( -30.24) >DroSim_CAF1 23486 117 - 1 CCCUCGAGGGCCAGUUCCAGGCACAUACUGGAUCCUCCAUCGGAUCCGGAGCCCUGCUGGGAAUUGCCGUCGUGGGAAUUAGUUCCUCCUCCACCCGAUCCCAGUCCCAAACUGGAC (((.(((.(((.((((((.((((....((((((((......)))))))).....)))).))))))))).))).)))........................(((((.....))))).. ( -48.00) >DroEre_CAF1 26730 117 - 1 CCUUCGAGGGCCAGUUCCAGGCACAUACUGGAUCCUCCGUCGGAUCCGGAGCCCUGCUGGGAGUUGCCGUCGUGGGAAUUGGCUCCUCCUCCGCCCGUUCCCAGUCCCAAACUGGAC .....(((((((((((((.(((((...((((((((......))))))))..(((....))).).))))......))))))))).))))............(((((.....))))).. ( -51.40) >DroYak_CAF1 24661 117 - 1 CCUUCGAGUGCCAGUUCCAGGCACAUACUGGAUCCUCCGUCGGAUCCGGAACCCUGCUGGGAGUUGCCGUCGUGUGAAUUGGCUCCUCCUCCGCCCGUUCCCAGUCCCAAGCUGGAC .....(((.(((((((((((((((...((((((((......))))))))..(((....))).).))))....)).))))))))..)))............(((((.....))))).. ( -44.20) >DroAna_CAF1 21547 108 - 1 CCCUCCAAGGCCAGCUCCAAGCACAUACUGGAGCCACCGUCGGAUC------CCUGCUGGGAGUUUCCAUCGUGCGAGUUG---CCACCGCCACCUCCUCCCAGACCCAGGCUGGAC ...((((.(((.(((((...((((....((((((..(((.(((...------.))).)))..).)))))..))))))))))---))...(((...((......))....))))))). ( -32.60) >consensus CCCUCGAGGGCCAGUUCCAGGCACAUACUGGAUCCUCCGUCGGAUCCGGA_CCCUGCUGGGAGUUGCCGUCGUGGGAAUUGGCUCCUCCUCCGCCCGAUCCCAGUCCCAAACUGGAC ....(((.(((.((((((.((((....((((((((......)))))))).....)))).))))))))).)))............................(((((.....))))).. (-23.23 = -26.20 + 2.97)


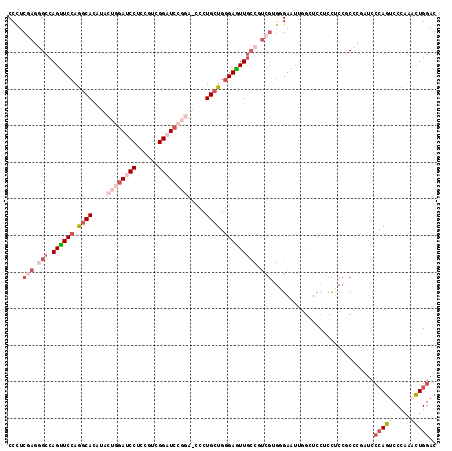
| Location | 23,408,921 – 23,409,038 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.73 |
| Mean single sequence MFE | -55.70 |
| Consensus MFE | -40.87 |
| Energy contribution | -39.85 |
| Covariance contribution | -1.02 |
| Combinations/Pair | 1.37 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539706 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23408921 117 + 27905053 GA---GGAUCCAGUAUGUGCCUGGAACUGGCCCUCGAGGGUGAACGCCUCUGCAAGGCGGGUGACUGCCGGGCGGGCGUGGCCUUCUUCCAGGCGGCCAUUCAGGCCGGAACCGAGGAUC ..---((.(((.......((((((((..((((..((.((((....))))((((..(((((....)))))..)))).)).))))...))))))))((((.....))))))).))....... ( -62.30) >DroVir_CAF1 45300 117 + 1 GC---AGUUCCAGCAUGUGUCUGGAGCUGGCUUUGGAGGGCGAACGUCUGUGCAAGGCAGGUGAUUGUCGGGCGGGCGUUGCCUUCUUCCAGGCAGCGAUACAGGCAGGCACCGACGAUC .(---(((((((((....).)))))))))((((((..((((....))))...))))))....(((((((((((..(((((((((......))))))).......))..)).))))))))) ( -52.81) >DroGri_CAF1 46161 117 + 1 GU---GGCUCAAGCAUGUGCCUGGAACUGGCGUUGGAGGGUGAGCGUCUCUGCAAAGCGGGCGAUUGUCGGGCGGGCGUUGCCUUCUUUCAGGCAGCCAUCCAAGCGGGCACCGACGAUC ..---.((((..((....(((.......))).(((.((((.......)))).))).))))))(((((((((((.(((..(((((......))))))))..((....)))).))))))))) ( -45.00) >DroYak_CAF1 24741 117 + 1 GA---GGAUCCAGUAUGUGCCUGGAACUGGCACUCGAAGGCGAACGCCUUUGCAAGGCGGGCGAUUGCCGGGCGGGCGUGGCCUUCUUCCAGGCGGCCAUUCAGGCCGGAACCGAGGAUC ..---((.(((.......((((((((.........((((((..((((((..((..(((((....)))))..)))))))).))))))))))))))((((.....))))))).))....... ( -57.60) >DroMoj_CAF1 44382 120 + 1 GCGGCGGCUCUAGCAUGUGUCUGGAGCUGGCCCUGGAAGGCGAACGCCUCUGCAAGGCAGGCGAUUGCCGAGCGGGCGUUGCCUUCUUCCAAGCAGCCAUCCAGGCGGGCACCGACGACC .(((((((((((((....).)))))))))((((.(((((((.(((((((..((..(((((....)))))..))))))))))))))))........(((.....))))))).)))...... ( -61.40) >DroAna_CAF1 21618 117 + 1 GU---GGCUCCAGUAUGUGCUUGGAGCUGGCCUUGGAGGGCGAACGGCUGUGCAAGGCGGGCGACUGUCGGGCGGGCGUGGCCUUCUUUCAAGCAGCCAUCCAGGCGGGCACCGAGGACC ..---(((((((((....)).)))))))(((((((((((((..(((.((((.(..(((((....)))))).)))).))).))))))......((.(((.....)))..))..))))).)) ( -55.10) >consensus GA___GGCUCCAGCAUGUGCCUGGAACUGGCCCUGGAGGGCGAACGCCUCUGCAAGGCGGGCGAUUGCCGGGCGGGCGUGGCCUUCUUCCAGGCAGCCAUCCAGGCGGGCACCGACGAUC .....((..((......(((((((((.........((((((..((((((..((..(((((....)))))..)))))))).)))))))))))))))(((.....))).))..))....... (-40.87 = -39.85 + -1.02)


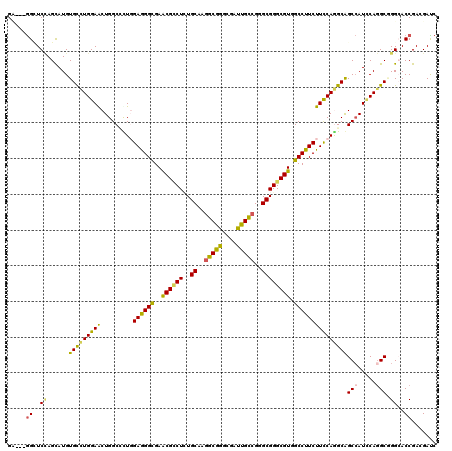
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:19:25 2006