
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,300,696 – 23,300,789 |
| Length | 93 |
| Max. P | 0.810688 |

| Location | 23,300,696 – 23,300,789 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 87.89 |
| Mean single sequence MFE | -26.14 |
| Consensus MFE | -21.20 |
| Energy contribution | -21.48 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562387 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
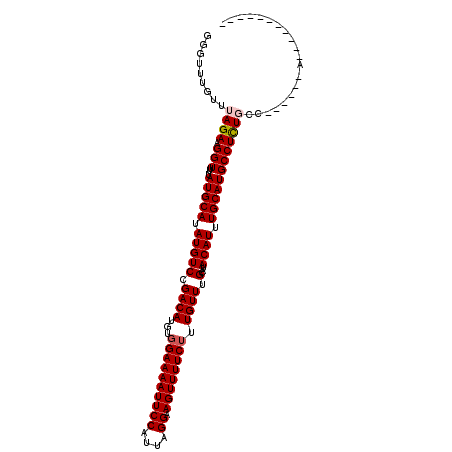
>3R_DroMel_CAF1 23300696 93 + 27905053 GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUCUGCC------------------ .........((((.(((...(((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).))))))))))))..------------------ ( -23.30) >DroPse_CAF1 103150 105 + 1 GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUUUCUCUCUGCCU------CUGCC ((((.......((((((...(((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).)))))))))))......))))------..... ( -27.12) >DroSec_CAF1 98143 95 + 1 GGGUUUGUUUAGAAGGU-UUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCAUUGUUUGCAUACAUUUGCAUGCCUCUGCC------AUC--------- .(((......(((.(((-..(((((.(((((.((((.(((((((.((((....)))).))))))))))).)...)))).))))))))))))))------...--------- ( -26.00) >DroSim_CAF1 102015 96 + 1 GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUCUGCC------AUC--------- .(((......(((.(((...(((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).))))))))))))))------...--------- ( -24.20) >DroYak_CAF1 108431 111 + 1 GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUCUGCCUCAGCCAAAACCUUCGCC ...........((((((((((((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).)))))......((....)).)))))))))... ( -29.10) >DroPer_CAF1 103586 105 + 1 GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUUUCUCUCUGCCU------CUGCC ((((.......((((((...(((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).)))))))))))......))))------..... ( -27.12) >consensus GGGUUUGUUUAGAAGGUUUUAUGCAUAUGUCCGACAUGUGGAAAAUUCCAUUAGGAAGUUUUCUUUGUUUGCAUACAUUUGCAUGCCUCUGCC______A___________ .........((((.(((...(((((.(((((.((((...((((((((((....)).)))))))).)))).)...)))).)))))))))))).................... (-21.20 = -21.48 + 0.28)


| Location | 23,300,696 – 23,300,789 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 87.89 |
| Mean single sequence MFE | -21.47 |
| Consensus MFE | -18.29 |
| Energy contribution | -18.07 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.810688 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23300696 93 - 27905053 ------------------GGCAGAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC ------------------...(((((.(((((.((((..(((......(((((.((((....)))))))))....)))...)))).))))).....))))).......... ( -18.60) >DroPse_CAF1 103150 105 - 1 GGCAG------AGGCAGAGAGAAAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC .(((.------.((((.........((((((....)))))).......(((((.((((....)))))))))....)))).......)))...................... ( -20.30) >DroSec_CAF1 98143 95 - 1 ---------GAU------GGCAGAGGCAUGCAAAUGUAUGCAAACAAUGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAA-ACCUUCUAAACAAACCC ---------..(------(..(((((.(((((.((((..(((......(((((.((((....)))))))))....)))...)))).)))))..-..)))))...))..... ( -19.60) >DroSim_CAF1 102015 96 - 1 ---------GAU------GGCAGAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC ---------..(------(..(((((.(((((.((((..(((......(((((.((((....)))))))))....)))...)))).))))).....)))))...))..... ( -18.90) >DroYak_CAF1 108431 111 - 1 GGCGAAGGUUUUGGCUGAGGCAGAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC ...((((((((((((((.((((...((((((....)))))).......(((((.((((....)))))))))....))))...))...)).))))))))))........... ( -31.10) >DroPer_CAF1 103586 105 - 1 GGCAG------AGGCAGAGAGAAAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC .(((.------.((((.........((((((....)))))).......(((((.((((....)))))))))....)))).......)))...................... ( -20.30) >consensus ___________U______GGCAGAGGCAUGCAAAUGUAUGCAAACAAAGAAAACUUCCUAAUGGAAUUUUCCACAUGUCGGACAUAUGCAUAAAACCUUCUAAACAAACCC ......................((((.(((((.((((..(((......(((((.((((....)))))))))....)))...)))).)))))....))))............ (-18.29 = -18.07 + -0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:18:00 2006