
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 23,112,644 – 23,112,804 |
| Length | 160 |
| Max. P | 0.939929 |

| Location | 23,112,644 – 23,112,764 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.50 |
| Mean single sequence MFE | -51.92 |
| Consensus MFE | -44.04 |
| Energy contribution | -44.12 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.939929 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
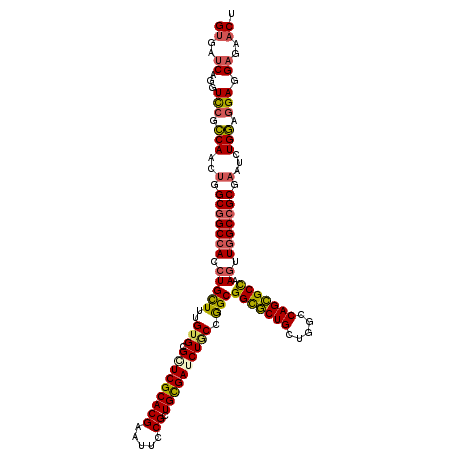
>3R_DroMel_CAF1 23112644 120 + 27905053 CGCUUCCGCCGACUCAGUUGGCAUUUGUUGCGGCACAAAUGUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGGUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUG .((..((((((((...)))((((...((((((((...........((((((((((....)))))..))))).(((((((....)))))))....)))))))).))))))))).))..... ( -52.40) >DroSim_CAF1 18508 120 + 1 CGCUUCCGCCGACUCAGUUGGCAUUUGUUGCGGCACAAAUGUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGCUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUG .((..((((((((...)))((((...((((((((...........((((((((((....)))))..))))).(((((((....)))))))....)))))))).))))))))).))..... ( -52.40) >DroEre_CAF1 16161 120 + 1 CGGUUCCGCCGACUCAGUUGGCGCUUGUUGCGGCACAAAUGUGAUCAGGUCCGCCAGCUGGCGGCCACCUGUUUUGUGCGGUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUG ((((.((((((((...)))(((((.....)).(((((((......((((((((((....)))))..))))).)))))))(((((((((.....)).))))))).)))))))).))))... ( -56.00) >DroYak_CAF1 16118 120 + 1 CGCUUCCGCCGACUCAGUUGGCGUUUGUUGCGGCACAAAUGUGAUCAAGUCCGCCAACUGGCGGCCACGUGCUUUGUGCGAUCGCACGAAUUCCGCUGCGAUCUACCGGCGGCGCUGCUG .((...(((((((...)))))))...(((((.(((((((.(((.....(.(((((....))))))....))))))))))(((((((((.....)).))))))).....)))))))..... ( -49.10) >DroAna_CAF1 15541 120 + 1 CGCUUCCGCCGUCUUAGCUGGCGGCUUUUGCGGCACAAGUGCGAGCAGGUUCGUCAACUAGCCGCCACCUGUUUUGUGCGCUCGCACGAGUUCCGUUGUGAUCUUCCAGCGGUACUGCUC (((....((((((......))))))....)))((....((((((((.((((.(....).))))((((.......)).)))))))))).(((.((((((.(.....))))))).))))).. ( -49.70) >consensus CGCUUCCGCCGACUCAGUUGGCGUUUGUUGCGGCACAAAUGUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGCUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUG .((...((((......(((((((...((((((((...........((((((((((....)))))..))))).(((((((....)))))))....)))))))).)))))))))))..)).. (-44.04 = -44.12 + 0.08)



| Location | 23,112,644 – 23,112,764 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.50 |
| Mean single sequence MFE | -49.90 |
| Consensus MFE | -40.12 |
| Energy contribution | -41.48 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23112644 120 - 27905053 CAGCAGCGCCGCCGGCAGAUCGCAGCGGAAUUCGUGCGACCGCACAAAGCAGGUGGCCGCCAGUUGGCGGACCUGAUCACAUUUGUGCCGCAACAAAUGCCAACUGAGUCGGCGGAAGCG .....((.((((((((.(.(((((.((.....))))))).)(((((((.(((((..(((((....))))))))))......))))))).(((.....))).......))))))))..)). ( -52.60) >DroSim_CAF1 18508 120 - 1 CAGCAGCGCCGCCGGCAGAUCGCAGCGGAAUUCGUGCGAGCGCACAAAGCAGGUGGCCGCCAGUUGGCGGACCUGAUCACAUUUGUGCCGCAACAAAUGCCAACUGAGUCGGCGGAAGCG .....((.((((((((...(((((.((.....)))))))(((((((((.(((((..(((((....))))))))))......)))))).)))................))))))))..)). ( -54.30) >DroEre_CAF1 16161 120 - 1 CAGCAGCGCCGCCGGCAGAUCGCAGCGGAAUUCGUGCGACCGCACAAAACAGGUGGCCGCCAGCUGGCGGACCUGAUCACAUUUGUGCCGCAACAAGCGCCAACUGAGUCGGCGGAACCG ........((((((((.(.(((((.((.....))))))).)(((((((.(((((..(((((....))))))))))......)))))))(((.....)))........))))))))..... ( -49.20) >DroYak_CAF1 16118 120 - 1 CAGCAGCGCCGCCGGUAGAUCGCAGCGGAAUUCGUGCGAUCGCACAAAGCACGUGGCCGCCAGUUGGCGGACUUGAUCACAUUUGUGCCGCAACAAACGCCAACUGAGUCGGCGGAAGCG .....((.((((((((.(((((((.((.....))))))))).(((.......))))))(((((((((((...(((..(((....)))...)))....))))))))).)).)))))..)). ( -51.10) >DroAna_CAF1 15541 120 - 1 GAGCAGUACCGCUGGAAGAUCACAACGGAACUCGUGCGAGCGCACAAAACAGGUGGCGGCUAGUUGACGAACCUGCUCGCACUUGUGCCGCAAAAGCCGCCAGCUAAGACGGCGGAAGCG ..((....((((((...................((((....)))).....((.((((((((.(((....))).(((..(((....))).)))..)))))))).))....))))))..)). ( -42.30) >consensus CAGCAGCGCCGCCGGCAGAUCGCAGCGGAAUUCGUGCGACCGCACAAAGCAGGUGGCCGCCAGUUGGCGGACCUGAUCACAUUUGUGCCGCAACAAACGCCAACUGAGUCGGCGGAAGCG ..((....((((((((.(.(((((.((.....))))))).)(((((((.(((((..(((((....))))))))))......)))))))...................))))))))..)). (-40.12 = -41.48 + 1.36)



| Location | 23,112,684 – 23,112,804 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.58 |
| Mean single sequence MFE | -52.60 |
| Consensus MFE | -44.34 |
| Energy contribution | -44.46 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537171 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 23112684 120 + 27905053 GUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGGUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUGGCCAGCGCCAAGUUGGCCGCGAAUCUGGAGGAGGAGAACU ((..((...(((.(((..(.(((((((...((((.(((((((((((((.....)).))))))).(((((((....)))))))..)))).))))))))))).)...))).))).))..)). ( -56.90) >DroSim_CAF1 18548 120 + 1 GUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGCUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUGGCCAGCGCCAAGUUGGCCGCGAAUCUGGAGGAGGAGAACU ....((.((.(((((....))))))).(((.((..(..((((((((((.....)).)))))...(((((((((((((.....)))))))..)))))).)))....)..)).))).))... ( -53.60) >DroEre_CAF1 16201 120 + 1 GUGAUCAGGUCCGCCAGCUGGCGGCCACCUGUUUUGUGCGGUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUGGCCAGCGCCAAGUUGGCCGCGAAUCUGGAGGAGGAGAACU ((..((...(((.((((.(.(((((((.((.....(((((((((((((.....)).))))))).(((((((....)))))))..))))..)).))))))).)..)))).))).))..)). ( -57.20) >DroYak_CAF1 16158 120 + 1 GUGAUCAAGUCCGCCAACUGGCGGCCACGUGCUUUGUGCGAUCGCACGAAUUCCGCUGCGAUCUACCGGCGGCGCUGCUGGCCAGCGCCAAGUUGGCCGCGAAUCUGGAGGAGGAGAACU ((..((...(((.(((..(.((((((((..(((..(((.(((((((((.....)).)))))))))).)))(((((((.....)))))))..).))))))).)...))).))).))..)). ( -55.10) >DroAna_CAF1 15581 120 + 1 GCGAGCAGGUUCGUCAACUAGCCGCCACCUGUUUUGUGCGCUCGCACGAGUUCCGUUGUGAUCUUCCAGCGGUACUGCUCUCCAGUGCUAAGUUAGCCGCUCAUCUGAGUGAUGAGAACU ((((((.((((.(....).))))((((.......)).))))))))..((((.((((((.(.....)))))))....))))....(.((((...))))).((((((.....)))))).... ( -40.20) >consensus GUGAUCAGGUCCGCCAACUGGCGGCCACCUGCUUUGUGCGCUCGCACGAAUUCCGCUGCGAUCUGCCGGCGGCGCUGCUGGCCAGCGCCAAGUUGGCCGCGAAUCUGGAGGAGGAGAACU ((..((...(((.(((..(.(((((((.(((((..(((.(((((((((.....)).)))))))))).)))(((((((.....))))))).)).))))))).)...))).))).))..)). (-44.34 = -44.46 + 0.12)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:16:10 2006