
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,620,540 – 22,620,659 |
| Length | 119 |
| Max. P | 0.951403 |

| Location | 22,620,540 – 22,620,630 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 81.47 |
| Mean single sequence MFE | -29.55 |
| Consensus MFE | -24.33 |
| Energy contribution | -24.17 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.787759 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22620540 90 + 27905053 CUGUUUU-----CCGCUGCGUGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUUAAAUCCGCUUCUGGAUAUACCCAGA ......(-----((((.(((((((((((.((....))))))))..)))))....)))))...(((((......))))).(((((......))))) ( -30.30) >DroPse_CAF1 76191 95 + 1 CUGUUGCUGCUGCUGCUGCGUGUUGCUCACACAUGCGGCGCAAAUUAUGUCAACGUGGAUUUCUGAAGUCAACUCCGCACUCGGGGAUAUCCGUG ..(((((...(((.(((((((((.......))))))))))))......).))))(((((....((....))..))))).(.((((....)))).) ( -27.40) >DroSec_CAF1 68801 90 + 1 CUGUUUU-----CCGCUGCGUGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAUCCCCAGA ....(((-----.(((..(((((.......)))))..))).)))..........(((((((((......))))))))).((((((....)))))) ( -33.00) >DroSim_CAF1 70592 90 + 1 CUGUUCU-----CCGCUGCGUGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAUCUCCAGA ......(-----((((.(((((((((((.((....))))))))..)))))....)))))...(((((......))))).((((((....)))))) ( -30.30) >DroYak_CAF1 71921 76 + 1 CUGCUUG-----CUGCUGCGUGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUC-------------- ....(((-----(.((..(((((.......)))))..))))))...........(((((((((......)))))))))...-------------- ( -26.70) >DroPer_CAF1 80367 92 + 1 CUGUUGCUGC---UGCUGCGUGUUGCUCACACAUGCGGCGCAAAUUAUGUCAACGUGGAUUUGUGAAGUCAACUCCGCACUCGGGGAUAUCCGUG ..((((((((---.(((((((((.......))))))))))))......).))))(((((.(((......))).))))).(.((((....)))).) ( -29.60) >consensus CUGUUUU_____CCGCUGCGUGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAUAUCCAGA .............((((((((((.......))))))))))..............(((((((((......)))))))))...((((....)))).. (-24.33 = -24.17 + -0.17)



| Location | 22,620,540 – 22,620,630 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 81.47 |
| Mean single sequence MFE | -27.91 |
| Consensus MFE | -18.65 |
| Energy contribution | -19.22 |
| Covariance contribution | 0.57 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802944 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22620540 90 - 27905053 UCUGGGUAUAUCCAGAAGCGGAUUUAACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACACGCAGCGG-----AAAACAG ((((((....))))))...((((((........)))))).(((((.......((((((((....)).))))))....))))).-----....... ( -27.10) >DroPse_CAF1 76191 95 - 1 CACGGAUAUCCCCGAGUGCGGAGUUGACUUCAGAAAUCCACGUUGACAUAAUUUGCGCCGCAUGUGUGAGCAACACGCAGCAGCAGCAGCAACAG ..(((......)))..((((((.(((....)))...)))..............(((((.((.((((((.....)))))))).)).)))))).... ( -26.80) >DroSec_CAF1 68801 90 - 1 UCUGGGGAUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACACGCAGCGG-----AAAACAG ((((((....))))))...(((((((......))))))).(((((.......((((((((....)).))))))....))))).-----....... ( -34.10) >DroSim_CAF1 70592 90 - 1 UCUGGAGAUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACACGCAGCGG-----AGAACAG (((((......)))))...(((((((......))))))).(((((.......((((((((....)).))))))....))))).-----....... ( -30.10) >DroYak_CAF1 71921 76 - 1 --------------GAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACACGCAGCAG-----CAAGCAG --------------...(((((((((......)))))))..((((.......((((((((....)).)))))).......)))-----)..)).. ( -25.44) >DroPer_CAF1 80367 92 - 1 CACGGAUAUCCCCGAGUGCGGAGUUGACUUCACAAAUCCACGUUGACAUAAUUUGCGCCGCAUGUGUGAGCAACACGCAGCA---GCAGCAACAG ..(((......)))..((((((.(((......))).))).................((.((.((((((.....)))))))).---)).))).... ( -23.90) >consensus UCUGGAUAUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACACGCAGCAG_____AAAACAG ..(((......)))...(.(((((((......))))))).)((((.......((((((((....)).))))))....)))).............. (-18.65 = -19.22 + 0.57)



| Location | 22,620,555 – 22,620,659 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 77.32 |
| Mean single sequence MFE | -32.90 |
| Consensus MFE | -21.83 |
| Energy contribution | -22.68 |
| Covariance contribution | 0.85 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.951403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22620555 104 + 27905053 UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUUAAAUCCGCUUCUGGAUAUACCCAGAUCACGCGCU--CGCAAUGGCUAGU-CCCCAUG-- .(((((((((....)))((((.............(((((((((......))))))))).(((((......)))))....)))).--)))))).......-.......-- ( -33.30) >DroSec_CAF1 68816 105 + 1 UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAUCCCCAGAUCACGCGCU--CACAAUGGCUAGUGCCCCCUG-- ......((((.((((((((((.............(((((((((......))))))))).((((((....))))))....)))).--))).)))....))))......-- ( -39.60) >DroSim_CAF1 70607 105 + 1 UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAUCUCCAGAUCACGCGCU--CACAAUGGCUAGUGCCCCAUG-- ......((((.((((((((((.............(((((((((......))))))))).((((((....))))))....)))).--))).)))....))))......-- ( -37.10) >DroEre_CAF1 76988 89 + 1 UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUUUGGGGAU------AUCACGCGCU--CACAAUGGCCA------------ ...((.((.((..((((((((.....(((((..((((((((((......)))))))))....)..)))------))...)))).--)))).))))))------------ ( -28.40) >DroYak_CAF1 71936 82 + 1 UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUC-------------------CACU--CACAAUGGCCCA----UCAAG-- ..((((((.((....)))))))).....((((..(((((...(((((......))))).))-------------------))).--.....))))...----.....-- ( -25.40) >DroPer_CAF1 80384 109 + 1 UGUUGCUCACACAUGCGGCGCAAAUUAUGUCAACGUGGAUUUGUGAAGUCAACUCCGCACUCGGGGAUAUCCGUGGCACACACACACACACGGACACAUGCCACAGACA .(((((........)))))........((((...((((...((((.......(((((....)))))...((((((.............))))))))))..)))).)))) ( -33.62) >consensus UGUUGCGCACACAUGUGGCGCAAAUUAUGUCAACGUGGAUUUGUGGAGUCAAAUCCGCUUCUGGGUAU__CCAGAUCACGCGCU__CACAAUGGCUAGU_CCCCAUG__ ..((((((.((....)))))))).....((((..(((((((((......)))))))))...(((......)))..................)))).............. (-21.83 = -22.68 + 0.85)



| Location | 22,620,555 – 22,620,659 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 77.32 |
| Mean single sequence MFE | -32.18 |
| Consensus MFE | -17.39 |
| Energy contribution | -18.48 |
| Covariance contribution | 1.10 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.54 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22620555 104 - 27905053 --CAUGGGG-ACUAGCCAUUGCG--AGCGCGUGAUCUGGGUAUAUCCAGAAGCGGAUUUAACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA --..(((..-.....)))(((((--.(((((((.((((((....)))))).(((((((....((.((........)).))...)))))))....))))))).))))).. ( -30.80) >DroSec_CAF1 68816 105 - 1 --CAGGGGGCACUAGCCAUUGUG--AGCGCGUGAUCUGGGGAUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA --.....(((....)))..(((.--((((.((..((((((....)))))).))(((((((......))))))).)))).)))....((((((((....)).)))))).. ( -41.40) >DroSim_CAF1 70607 105 - 1 --CAUGGGGCACUAGCCAUUGUG--AGCGCGUGAUCUGGAGAUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA --.....(((....)))..(((.--((((.((..(((((......))))).))(((((((......))))))).)))).)))....((((((((....)).)))))).. ( -37.40) >DroEre_CAF1 76988 89 - 1 ------------UGGCCAUUGUG--AGCGCGUGAU------AUCCCCAAAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA ------------.......(((.--(((((.((..------.....))...))(((((((......)))))))..))).)))....((((((((....)).)))))).. ( -25.70) >DroYak_CAF1 71936 82 - 1 --CUUGA----UGGGCCAUUGUG--AGUG-------------------GAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA --.(..(----((..((((....--.)))-------------------)....(((((((......))))))).)))..)......((((((((....)).)))))).. ( -24.40) >DroPer_CAF1 80384 109 - 1 UGUCUGUGGCAUGUGUCCGUGUGUGUGUGUGUGCCACGGAUAUCCCCGAGUGCGGAGUUGACUUCACAAAUCCACGUUGACAUAAUUUGCGCCGCAUGUGUGAGCAACA (((((((((((..((..((....))..))..)))))))))))...(((....))).((((.((.((((......((.((.((.....)))).))....)))))))))). ( -33.40) >consensus __CAUGGGG_ACUAGCCAUUGUG__AGCGCGUGAUCUGG__AUACCCAGAAGCGGAUUUGACUCCACAAAUCCACGUUGACAUAAUUUGCGCCACAUGUGUGCGCAACA .................((((((...(((.......(((......))).....(((((((......))))))).)))...))))))((((((((....)).)))))).. (-17.39 = -18.48 + 1.10)

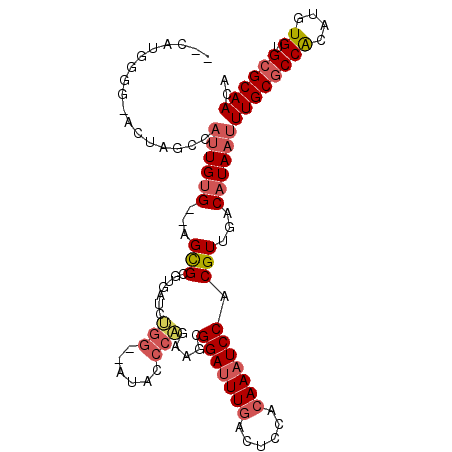

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:12:18 2006