
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,570,344 – 22,570,474 |
| Length | 130 |
| Max. P | 0.966566 |

| Location | 22,570,344 – 22,570,438 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.23 |
| Mean single sequence MFE | -21.56 |
| Consensus MFE | -14.09 |
| Energy contribution | -13.98 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22570344 94 - 27905053 AAAUCACUGGCUGUUACUCAAGCAGUCAUGUGAUUUGAUUGAUUCGCGCUAAAAAUCCGUAGUCAAACAAAC-G-CCAGAUCCAGCACU---------CGCAGA---------------A (((((((((((((((.....)))))))).)))))))..((((((.(((.........)))))))))......-(-(.((........))---------.))...---------------. ( -22.70) >DroVir_CAF1 25268 98 - 1 AAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCUGCACUCAAACACUU-GGGCAG-UCAUACAUC-----CCUACGAUGU---------------G (((((((((((((((.....)))))))).)))))))(((((......((.........)).(((((....))-))))))-)).((((((-----.....)))))---------------) ( -28.60) >DroGri_CAF1 31733 99 - 1 AAAUCACUGGUUGUUAGUCAAGUAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCCCACUCAAACACAAAAUACAG-UCCAACUUG-----UCUAUGAUGA---------------A .......(.((..(.((.(((((.(...((((.((((.((((....................))))...)))).)))).-..).)))))-----.)))..)).)---------------. ( -12.15) >DroWil_CAF1 55590 108 - 1 AAAUCAGUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCGUACUCAAACAAGC-G-ACAGCCCC-GCACA---------AAAGCGCACUCAAACACACACA ......(((((((((.....)))))))))(((.(((((.((.....((((........((......))..((-(-.......)-))...---------..)))))).))))))))..... ( -23.70) >DroMoj_CAF1 21608 94 - 1 AAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCUGCACUCAAACACUU-GCACAG-UGAAGCAUC-----UCUAUGG------------------- (((((((((((((((.....)))))))).)))))))(((...(((((((.........))...(((....))-)....)-))))..)))-----.......------------------- ( -22.80) >DroAna_CAF1 20184 102 - 1 AAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCCUUCUCAAACAAAC-A-GC-GACCCAACCCCCACGUUCUCCGCUCC---------------A (((((((((((((((.....)))))))).)))))))....................................-(-((-(....(((......)))...))))..---------------. ( -19.40) >consensus AAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCGCACUCAAACAAAC_G_ACAG_UCCAACACC_____UCUACGAUGA_______________A (((((((((((((((.....)))))))).))))))).................................................................................... (-14.09 = -13.98 + -0.11)


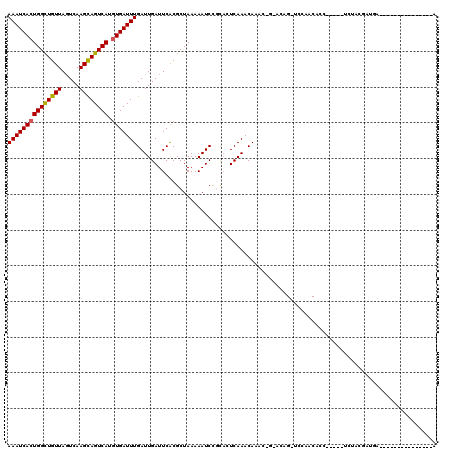
| Location | 22,570,360 – 22,570,474 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.74 |
| Mean single sequence MFE | -32.00 |
| Consensus MFE | -22.45 |
| Energy contribution | -22.98 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.781317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22570360 114 + 27905053 UCUGG-C-GUUUGUUUGACUACGGAUUUUUAGCGCGAAUCAAUCAAAUCACAUGACUGCUUGAGUAACAGCCAGUGAUUUGAUUGAUGAACUGCGUCAACAGCGA----GAGGGGCGAGC (((.(-(-((((((......)))))).....(((((.((((((((((((((.((.(((.........))).))))))))))))))))....))))).....)).)----))......... ( -35.20) >DroVir_CAF1 25288 109 + 1 -CUGCCC-AAGUGUUUGAGUGCAGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUGCUUGACUAACAGCCAGUGAUUUGAUUGAUGAACUGCGUUGCCAGAUA---GCUGUG------ -((((((-((....))).).)))).......((((..((((((((((((((.((.(((.........))).))))))))))))))))..)).)).....(((...---.)))..------ ( -31.70) >DroGri_CAF1 31753 113 + 1 -CUGUAUUUUGUGUUUGAGUGGGGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUACUUGACUAACAACCAGUGAUUUGAUUGAUGAACUGUUUUGUCAGAUAGUAGCUGUG------ -((.(((((.......))))).)).....((((....((((((((((((((.((.................))))))))))))))))..((((((......)))))).))))..------ ( -27.63) >DroWil_CAF1 55620 108 + 1 GCUGU-C-GCUUGUUUGAGUACGGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUGCUUGACUAACAGCCACUGAUUUGAUUGAUGAACUGCGUCAACGUCAA----UGUGG------ .((((-.-((((....)))))))).......((((..(((((((((((((..((.(((.........))).)).)))))))))))))..)).))...........----.....------ ( -27.00) >DroMoj_CAF1 21624 109 + 1 -CUGUGC-AAGUGUUUGAGUGCAGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUGCUUGACUAACAGCCAGUGAUUUGAUUGAUGAACUGCGUUGCCGGAUA---GCUCUG------ -(((.((-((..(((((....))))).....((((..((((((((((((((.((.(((.........))).))))))))))))))))..)).)).)))))))...---......------ ( -33.30) >DroAna_CAF1 20209 110 + 1 UC-GC-U-GUUUGUUUGAGAAGGGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUGCUUGACUAACAGCCAGUGAUUUGAUUGAUGAACUGCGUCAACGGCGAC---GACGAGU---- ((-((-(-(((....((((((....))))))((((..((((((((((((((.((.(((.........))).))))))))))))))))..)).))...)))))))).---.......---- ( -37.20) >consensus _CUGU_C_GUGUGUUUGAGUACGGAUUUUUAGCGUGAAUCAAUCAAAUCACAUGACUGCUUGACUAACAGCCAGUGAUUUGAUUGAUGAACUGCGUCAACAGAUA____CUGGG______ ...............................((((..((((((((((((((.((.(((.........))).))))))))))))))))..)).)).......................... (-22.45 = -22.98 + 0.53)
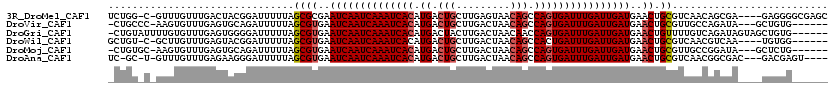

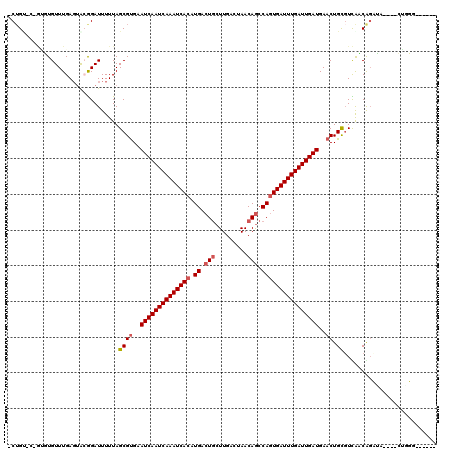
| Location | 22,570,360 – 22,570,474 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.74 |
| Mean single sequence MFE | -34.24 |
| Consensus MFE | -28.01 |
| Energy contribution | -27.93 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -4.25 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966566 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22570360 114 - 27905053 GCUCGCCCCUC----UCGCUGUUGACGCAGUUCAUCAAUCAAAUCACUGGCUGUUACUCAAGCAGUCAUGUGAUUUGAUUGAUUCGCGCUAAAAAUCCGUAGUCAAACAAAC-G-CCAGA .........((----(.(((((((((.......((((((((((((((((((((((.....)))))))).))))))))))))))..(((.........))).))).))))...-)-).))) ( -36.00) >DroVir_CAF1 25288 109 - 1 ------CACAGC---UAUCUGGCAACGCAGUUCAUCAAUCAAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCUGCACUCAAACACUU-GGGCAG- ------....((---(....)))...((.((..((((((((((((((((((((((.....)))))))).))))))))))))))..)))).......((((.(.(((....))-))))))- ( -38.10) >DroGri_CAF1 31753 113 - 1 ------CACAGCUACUAUCUGACAAAACAGUUCAUCAAUCAAAUCACUGGUUGUUAGUCAAGUAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCCCACUCAAACACAAAAUACAG- ------...(((......(((......)))...((((((((((((((((..((((.....))))..)).))))))))))))))....))).............................- ( -22.80) >DroWil_CAF1 55620 108 - 1 ------CCACA----UUGACGUUGACGCAGUUCAUCAAUCAAAUCAGUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCGUACUCAAACAAGC-G-ACAGC ------.....----.....((((.(((.((..((((((((((((((((((((((.....))))))))).)))))))))))))..))...........((......))..))-)-.)))) ( -34.40) >DroMoj_CAF1 21624 109 - 1 ------CAGAGC---UAUCCGGCAACGCAGUUCAUCAAUCAAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCUGCACUCAAACACUU-GCACAG- ------..((((---(...((....)).)))))((((((((((((((((((((((.....)))))))).))))))))))))))..............((((..........)-)))...- ( -37.40) >DroAna_CAF1 20209 110 - 1 ----ACUCGUC---GUCGCCGUUGACGCAGUUCAUCAAUCAAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCCUUCUCAAACAAAC-A-GC-GA ----(((((((---(.......))))).)))..((((((((((((((((((((((.....)))))))).))))))))))))))...((((......................-)-))-). ( -36.75) >consensus ______CACAG____UAGCCGUCAACGCAGUUCAUCAAUCAAAUCACUGGCUGUUAGUCAAGCAGUCAUGUGAUUUGAUUGAUUCACGCUAAAAAUCCGCACUCAAACAAAC_G_ACAG_ ..........................((.((..((((((((((((((((((((((.....)))))))).))))))))))))))..))))............................... (-28.01 = -27.93 + -0.08)


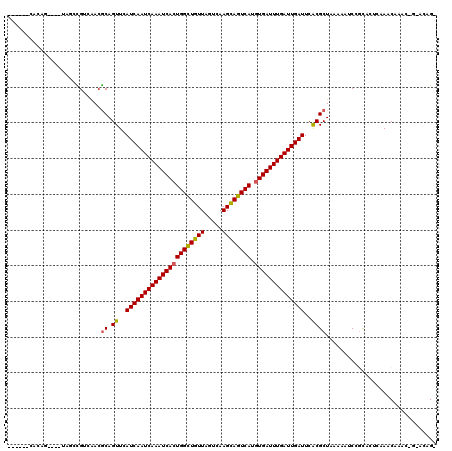
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:11:59 2006