
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,554,228 – 22,554,321 |
| Length | 93 |
| Max. P | 0.966225 |

| Location | 22,554,228 – 22,554,321 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 95.16 |
| Mean single sequence MFE | -29.55 |
| Consensus MFE | -27.50 |
| Energy contribution | -27.38 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.966225 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22554228 93 + 27905053 CCCAACUGGCCAGCUCCUAUCCAGCUCGGCUCGAGCGGAAGAAACAGCAAUCGGGCAAACAAACAGCGCUAUAAAAUGCGAAAACAGAGUGGG ((((.((((((.(((.((.(((.(((((...))))))))))....))).....)))..........(((........)))....)))..)))) ( -26.30) >DroSec_CAF1 2461 93 + 1 CCCACCUGGCCAGCUCCUAUCCAGCUCGUCUCGAGCGGAAGAAACAGCAAUUGGGCAAACAAACAGCGCCAUAAAAUGCGAAAGCAGAGUGGG (((((.(((((.(((.((.(((.(((((...))))))))))....)))..(((......)))...).)))).....(((....)))..))))) ( -30.90) >DroSim_CAF1 2422 93 + 1 CCCACCUGGCCAGCUCCUAUCCAGCUCGUCUCGAGCGGAAGAAACAGCAAUUGGGCAAACAAACAGCGCCAUAAAAUGCGAGAACAGAGUGGG ((((((((.....(((...(((.(((((...)))))))).......(((.((.(((...........)))...)).))))))..))).))))) ( -30.60) >DroYak_CAF1 2493 93 + 1 CCCACCUGGCCAGUUCCUAUCCAGCUCGGCUCGAGCGGAAGAAACAACAAUUGGGCAAACAAACAGCGCCAUAAAAUGCGAGAGCAGAGUGGG (((((.(((((.(((.((.(((.(((((...)))))))))).))).....(((......)))...).)))).....(((....)))..))))) ( -30.40) >consensus CCCACCUGGCCAGCUCCUAUCCAGCUCGGCUCGAGCGGAAGAAACAGCAAUUGGGCAAACAAACAGCGCCAUAAAAUGCGAAAACAGAGUGGG (((((((((((.(((.((.(((.(((((...))))))))))....))).....)))..........(((........)))....))).))))) (-27.50 = -27.38 + -0.12)


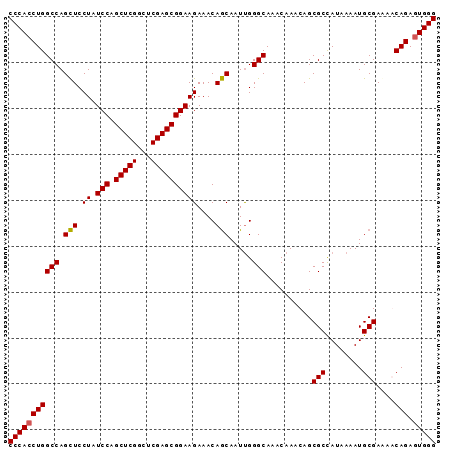
| Location | 22,554,228 – 22,554,321 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 95.16 |
| Mean single sequence MFE | -32.27 |
| Consensus MFE | -29.91 |
| Energy contribution | -29.85 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.723391 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22554228 93 - 27905053 CCCACUCUGUUUUCGCAUUUUAUAGCGCUGUUUGUUUGCCCGAUUGCUGUUUCUUCCGCUCGAGCCGAGCUGGAUAGGAGCUGGCCAGUUGGG .............(((........)))...........((((((((((((((((((((((((...))))).))).)))))).)).)))))))) ( -29.10) >DroSec_CAF1 2461 93 - 1 CCCACUCUGCUUUCGCAUUUUAUGGCGCUGUUUGUUUGCCCAAUUGCUGUUUCUUCCGCUCGAGACGAGCUGGAUAGGAGCUGGCCAGGUGGG ((((((..(((...(((.....(((.((.........)))))..))).((((((((((((((...))))).))).)))))).)))..)))))) ( -33.80) >DroSim_CAF1 2422 93 - 1 CCCACUCUGUUCUCGCAUUUUAUGGCGCUGUUUGUUUGCCCAAUUGCUGUUUCUUCCGCUCGAGACGAGCUGGAUAGGAGCUGGCCAGGUGGG (((((.(((..(..(((.....(((.((.........)))))..))).((((((((((((((...))))).))).)))))).)..)))))))) ( -34.40) >DroYak_CAF1 2493 93 - 1 CCCACUCUGCUCUCGCAUUUUAUGGCGCUGUUUGUUUGCCCAAUUGUUGUUUCUUCCGCUCGAGCCGAGCUGGAUAGGAACUGGCCAGGUGGG ((((((..(((...........(((.((.........)))))......((((((((((((((...))))).))).)))))).)))..)))))) ( -31.80) >consensus CCCACUCUGCUCUCGCAUUUUAUGGCGCUGUUUGUUUGCCCAAUUGCUGUUUCUUCCGCUCGAGACGAGCUGGAUAGGAGCUGGCCAGGUGGG (((((.(((.............(((.((.........)))))...(((((((((((((((((...))))).))).)))))).))))))))))) (-29.91 = -29.85 + -0.06)


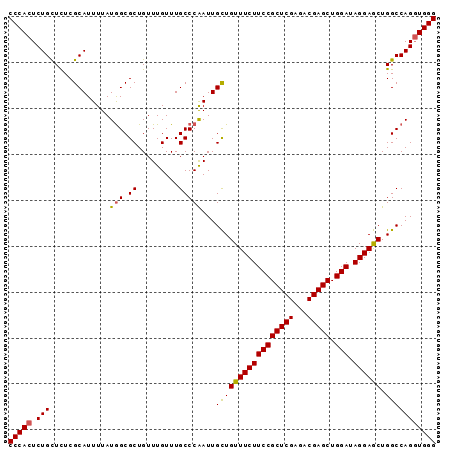
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:11:47 2006