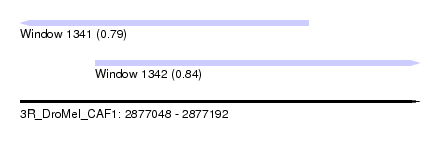

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,877,048 – 2,877,192 |
| Length | 144 |
| Max. P | 0.844151 |

| Location | 2,877,048 – 2,877,152 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.12 |
| Mean single sequence MFE | -24.07 |
| Consensus MFE | -20.29 |
| Energy contribution | -20.10 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.793715 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
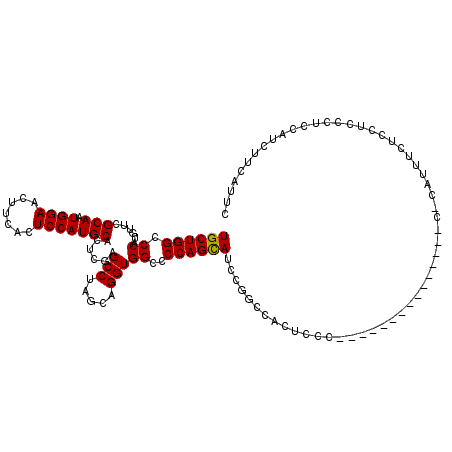
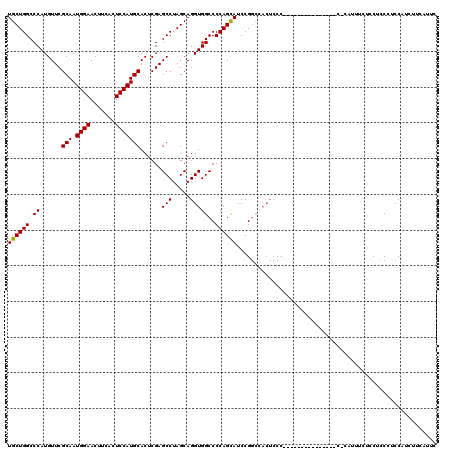
>3R_DroMel_CAF1 2877048 104 - 27905053 UGCUGGCCCAUGUUCGCAAUGGAACUUCACUCCAUGCACUCGAGCCUAGCAGGUGGCCCCAGUAUCCGGCCACUCCC---------------C-AUUUUCUCCUCCCUCCUUCUUCGUUC (((((((.(..((..(((.((((.......)))))))))..).))).))))(((((((.........)))))))...---------------.-.......................... ( -26.70) >DroSec_CAF1 8191 92 - 1 UGCUGGCCCAUGUUCGCAAUGGAACUUCACUCCAUGCACUCGAGCCUAGCAGGUGGCCCCAGCAUCCG----------------------------UUUCUCCUCCCUCCAUCUUCAUUC ((((((.((......(((.((((.......)))))))......(((.....)))))..))))))....----------------------------........................ ( -20.60) >DroSim_CAF1 10590 105 - 1 UGCUGGCCCAUGUUCGCAAUGGAACUUCACUCCAUGCACUCGAGCCUAGCAGGUGGCCCCAGCAUCCGGCCACUCCC---------------CCCAUUUCUCCUCCCUCCAUCUUCAUUC (((((((.(..((..(((.((((.......)))))))))..).))).))))(((((((.........)))))))...---------------............................ ( -26.70) >DroEre_CAF1 9209 120 - 1 UGCUGGCCCAUGUUCGCAAUGGAACUUCACUCCAUGCACUCGAGCCUAGCAGGUGGCCCCAGCAUCCGGCGACUCCCCUAAUCCUCCAUUCCCCCAUUCCUGCUCCCUCCUUCUUCAUUC (((((((.(..((..(((.((((.......)))))))))..).))).))))(((.(((.........))).))).............................................. ( -22.30) >consensus UGCUGGCCCAUGUUCGCAAUGGAACUUCACUCCAUGCACUCGAGCCUAGCAGGUGGCCCCAGCAUCCGGCCACUCCC_______________C_CAUUUCUCCUCCCUCCAUCUUCAUUC ((((((.((......(((.((((.......)))))))......(((.....)))))..))))))........................................................ (-20.29 = -20.10 + -0.19)

| Location | 2,877,075 – 2,877,192 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.81 |
| Mean single sequence MFE | -40.98 |
| Consensus MFE | -38.35 |
| Energy contribution | -38.35 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.844151 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
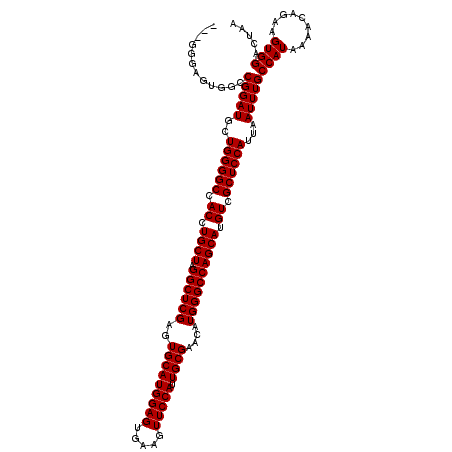
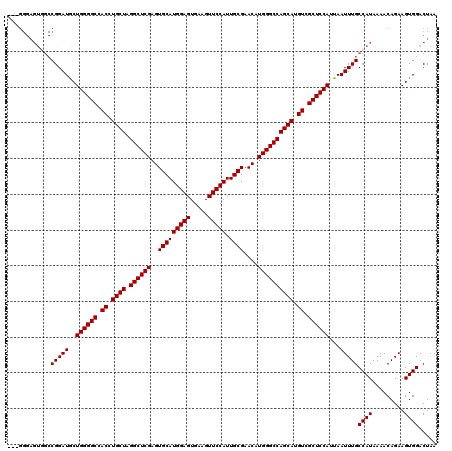
>3R_DroMel_CAF1 2877075 117 + 27905053 ---GGGAGUGGCCGGAUACUGGGGCCACCUGCUAGGCUCGAGUGCAUGGAGUGAAGUUCCAUUGCGAACAUGGGCCAGCAUGUCGCUCCAUUAAUUUGCCAUAAAACAGAAGUGGACUAA ---....(((((..(.((.((((((.((.((((.((((((..(((((((((.....))))).))))....)))))))))).)).)))))).)).)..))))).................. ( -42.40) >DroSec_CAF1 8215 108 + 1 ------------CGGAUGCUGGGGCCACCUGCUAGGCUCGAGUGCAUGGAGUGAAGUUCCAUUGCGAACAUGGGCCAGCAUGUCGCUCCAUUAAUUUGCCAUAAAACAGAAGUGGACUUA ------------(((((..((((((.((.((((.((((((..(((((((((.....))))).))))....)))))))))).)).))))))...)))))((((.........))))..... ( -38.40) >DroSim_CAF1 10618 117 + 1 ---GGGAGUGGCCGGAUGCUGGGGCCACCUGCUAGGCUCGAGUGCAUGGAGUGAAGUUCCAUUGCGAACAUGGGCCAGCAUGUCGCUCCAUUAAUUUGCCAUAAAACAGAAGUGGACUAA ---....(((((.((((..((((((.((.((((.((((((..(((((((((.....))))).))))....)))))))))).)).))))))...))))))))).................. ( -42.70) >DroEre_CAF1 9249 120 + 1 UAGGGGAGUCGCCGGAUGCUGGGGCCACCUGCUAGGCUCGAGUGCAUGGAGUGAAGUUCCAUUGCGAACAUGGGCCAGCAUGUCGCUCCAUUAAUUUGCCAUAAAACAGAAGUGGACUAA ......(((((.(((((..((((((.((.((((.((((((..(((((((((.....))))).))))....)))))))))).)).))))))...))))).).....((....)).)))).. ( -40.40) >consensus ___GGGAGUGGCCGGAUGCUGGGGCCACCUGCUAGGCUCGAGUGCAUGGAGUGAAGUUCCAUUGCGAACAUGGGCCAGCAUGUCGCUCCAUUAAUUUGCCAUAAAACAGAAGUGGACUAA ............(((((..((((((.((.((((.((((((..(((((((((.....))))).))))....)))))))))).)).))))))...)))))((((.........))))..... (-38.35 = -38.35 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:32 2006