| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,874,183 – 2,874,301 |
| Length | 118 |
| Max. P | 0.849161 |

| Location | 2,874,183 – 2,874,301 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.60 |
| Mean single sequence MFE | -41.45 |
| Consensus MFE | -33.01 |
| Energy contribution | -33.33 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.706374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
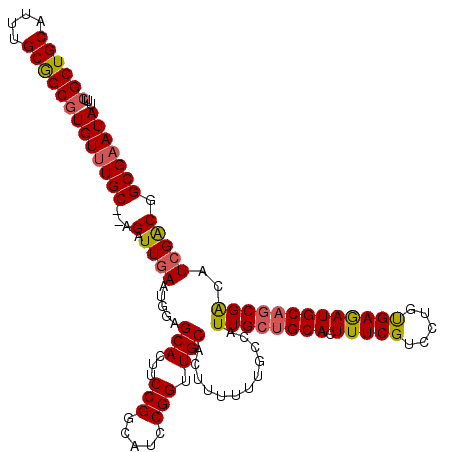
>3R_DroMel_CAF1 2874183 118 + 27905053 UUUGGCUGCAUUUGCGGCGUGUUUGC--AGAUUGAAUGGAGCACUUCCGGCAUCCGGUUGCACUUUUUUGCCAUUGCUGCACUUUCGUCCUGUGAAAUGCAGCGACAUCGACGGCGAAUA ....(((((....))))).(((((((--...((((.((..(((...((((...)))).)))............((((((((.(((((.....)))))))))))))))))))..))))))) ( -38.90) >DroSec_CAF1 5323 118 + 1 UUUCGCUGCAUUUGCAGCGUGUUUGC--AGAUUGAAUGGAGCACUUCCGGAAUCCGGUUGCACUUUUUUGCCAUUGCUGCACUUUCGUCCUGUGAGAUGCAGCGACAUCGACGGCGAAUA ...((((((....))))))(((((((--...((((.((..(((...((((...)))).)))............((((((((.(((((.....)))))))))))))))))))..))))))) ( -41.10) >DroSim_CAF1 5676 120 + 1 UUUCGCUGCAUUUGCGGCGUGUUUGCUCGGAUUGAAUGGAGCACUUCCGGCAUCCGGUUGCACUUUUUUUCCAUUGGAGCACUUUCGUCCUGUGAGAUGCAGCGACAUGGACGGCGAAUA ..(((((((((((((((((....(((((.(((.(((.((.(((...((((...)))).))).))....))).))).)))))....)).)).)).)))))))))))............... ( -42.50) >DroEre_CAF1 6276 117 + 1 UUUCGCUGCGUUUGCGGCGUGUUUGC--AGAUUGAAUGGAGCACUUCCGACAUCCGGUUGCACUUU-UUGCCAUUGCUGCACUUUCGUCCUGCGAGAUGCAGCGGCUUCGGCUGCGUAUA ...((((((....))))))..(((((--((..((((((.((((...(((.....)))..(((....-.)))...)))).))..))))..)))))))((((.(((((....))))))))). ( -43.30) >consensus UUUCGCUGCAUUUGCGGCGUGUUUGC__AGAUUGAAUGGAGCACUUCCGGCAUCCGGUUGCACUUUUUUGCCAUUGCUGCACUUUCGUCCUGUGAGAUGCAGCGACAUCGACGGCGAAUA ...((((((....))))))(((((((...(.((((.....(((...(((.....))).)))............((((((((.(((((.....)))))))))))))..))))).))))))) (-33.01 = -33.33 + 0.31)

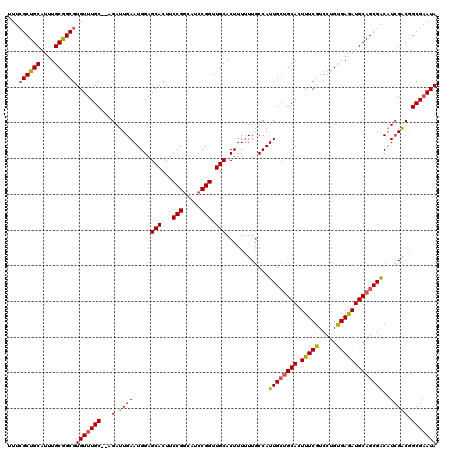
| Location | 2,874,183 – 2,874,301 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.60 |
| Mean single sequence MFE | -37.45 |
| Consensus MFE | -28.96 |
| Energy contribution | -29.77 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
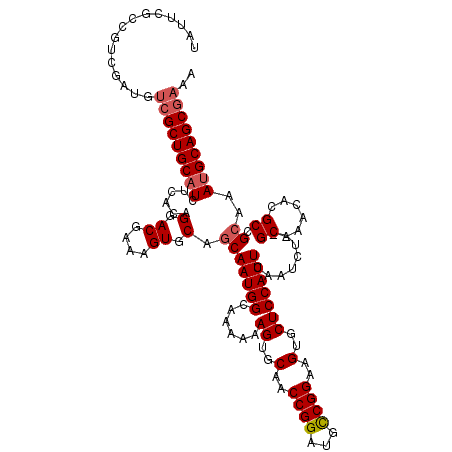
>3R_DroMel_CAF1 2874183 118 - 27905053 UAUUCGCCGUCGAUGUCGCUGCAUUUCACAGGACGAAAGUGCAGCAAUGGCAAAAAAGUGCAACCGGAUGCCGGAAGUGCUCCAUUCAAUCU--GCAAACACGCCGCAAAUGCAGCCAAA .................(((((((((....((.((....(((((.(((((......((..(..((((...))))..)..)))))))....))--)))....))))..))))))))).... ( -35.90) >DroSec_CAF1 5323 118 - 1 UAUUCGCCGUCGAUGUCGCUGCAUCUCACAGGACGAAAGUGCAGCAAUGGCAAAAAAGUGCAACCGGAUUCCGGAAGUGCUCCAUUCAAUCU--GCAAACACGCUGCAAAUGCAGCGAAA ..((((((((.(((((....)))))..)).)).))))..(((((.(((((......((..(..((((...))))..)..)))))))....))--)))....((((((....))))))... ( -39.50) >DroSim_CAF1 5676 120 - 1 UAUUCGCCGUCCAUGUCGCUGCAUCUCACAGGACGAAAGUGCUCCAAUGGAAAAAAAGUGCAACCGGAUGCCGGAAGUGCUCCAUUCAAUCCGAGCAAACACGCCGCAAAUGCAGCGAAA ...............(((((((((......((.((....(((((.....(((....((..(..((((...))))..)..))...))).....)))))....))))....))))))))).. ( -37.40) >DroEre_CAF1 6276 117 - 1 UAUACGCAGCCGAAGCCGCUGCAUCUCGCAGGACGAAAGUGCAGCAAUGGCAA-AAAGUGCAACCGGAUGUCGGAAGUGCUCCAUUCAAUCU--GCAAACACGCCGCAAACGCAGCGAAA .....((((..(((((((((((((.(((.....)))..)))))))...)))..-..((..(..(((.....)))..)..))...)))...))--)).....(((.((....)).)))... ( -37.00) >consensus UAUUCGCCGUCGAUGUCGCUGCAUCUCACAGGACGAAAGUGCAGCAAUGGCAAAAAAGUGCAACCGGAUGCCGGAAGUGCUCCAUUCAAUCU__GCAAACACGCCGCAAAUGCAGCGAAA ...............(((((((((......(.((....)).).(((((((......((..(..((((...))))..)..)))))))........((......)).))..))))))))).. (-28.96 = -29.77 + 0.81)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:27 2006