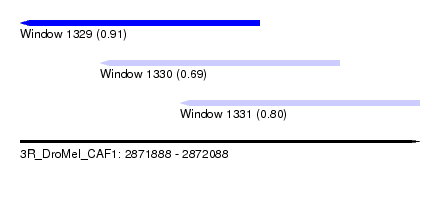

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,871,888 – 2,872,088 |
| Length | 200 |
| Max. P | 0.905110 |

| Location | 2,871,888 – 2,872,008 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.17 |
| Mean single sequence MFE | -34.65 |
| Consensus MFE | -34.65 |
| Energy contribution | -34.65 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.905110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
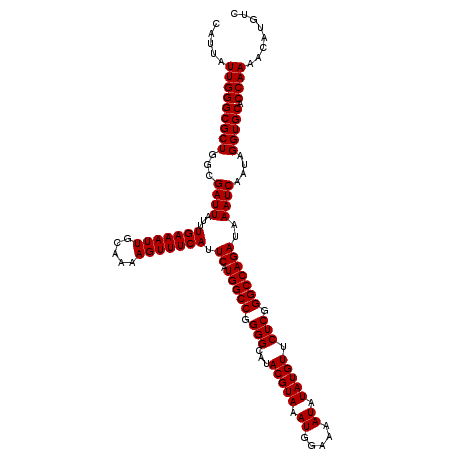
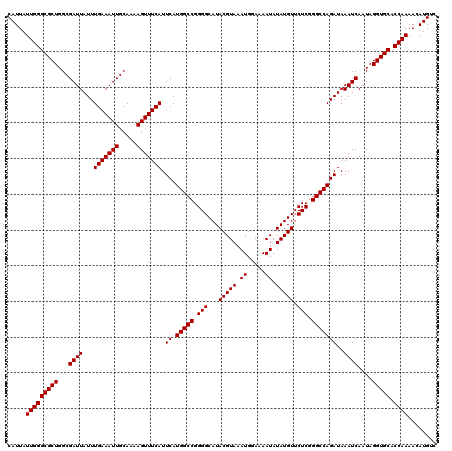
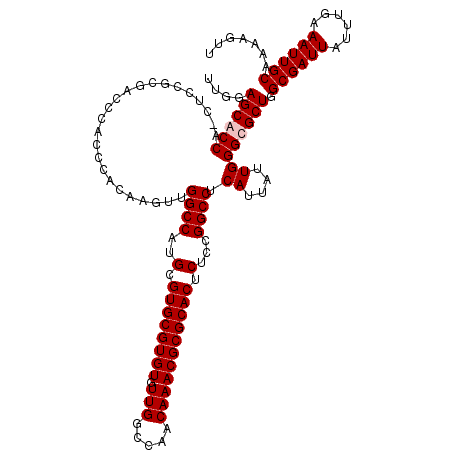
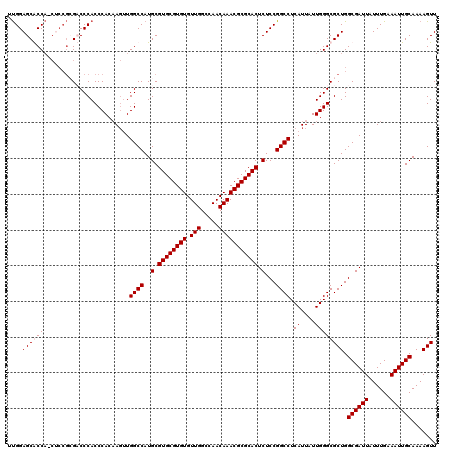
>3R_DroMel_CAF1 2871888 120 - 27905053 CAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGUUCUCGGGCCAGAUAAAUCAAUAGGUGCACCAAAACAUGCC .....(((((((((...((((...(((((((.....))))))).((.(((((.(((...(((((.((.....)).))))).))).)))))))..))))....))))).))))........ ( -35.10) >DroSec_CAF1 3082 120 - 1 CAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGUUCUCGGGCCAGAUAAAUCAAUAGGUGCACCAAAACAUGUC .....(((((((((...((((...(((((((.....))))))).((.(((((.(((...(((((.((.....)).))))).))).)))))))..))))....))))).))))........ ( -35.10) >DroSim_CAF1 3457 120 - 1 CAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGUUCUCGGGCCAGAUAAAUCAAUAGGUGCACCAAAACAUGUC .....(((((((((...((((...(((((((.....))))))).((.(((((.(((...(((((.((.....)).))))).))).)))))))..))))....))))).))))........ ( -35.10) >DroEre_CAF1 4027 120 - 1 CAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGUUCUCAGGCCAGAUAAAUCAAUAGGUGCACCAAAACAUGUC .....(((((((((...((((...(((((((.....))))))).((.(((((.(((...(((((.((.....)).))))).))).)))))))..))))....))))).))))........ ( -33.30) >consensus CAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGUUCUCGGGCCAGAUAAAUCAAUAGGUGCACCAAAACAUGUC .....(((((((((...((((...(((((((.....))))))).((.(((((.(((...(((((.((.....)).))))).))).)))))))..))))....))))).))))........ (-34.65 = -34.65 + -0.00)

| Location | 2,871,928 – 2,872,048 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -40.00 |
| Consensus MFE | -40.00 |
| Energy contribution | -40.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.03 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685145 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
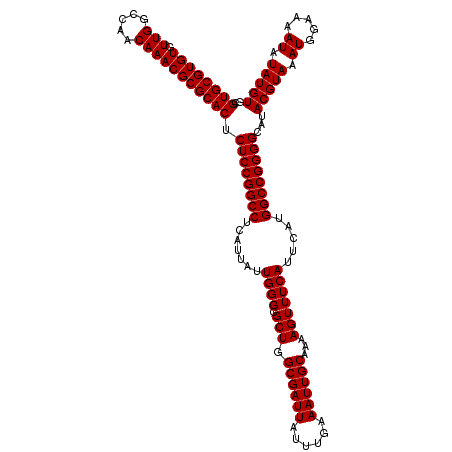
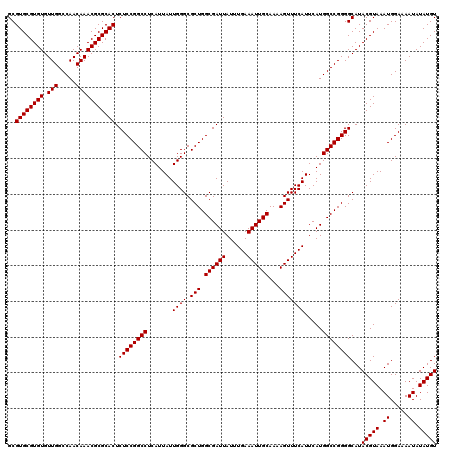
>3R_DroMel_CAF1 2871928 120 - 27905053 GCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGU ..((((((((.(((.....))))))))))).((((((((.......((((.(((.((((((......))))))...))))))).....))))))))...(((((.((.....)).))))) ( -40.00) >DroSec_CAF1 3122 120 - 1 GCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGU ..((((((((.(((.....))))))))))).((((((((.......((((.(((.((((((......))))))...))))))).....))))))))...(((((.((.....)).))))) ( -40.00) >DroSim_CAF1 3497 120 - 1 GCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGU ..((((((((.(((.....))))))))))).((((((((.......((((.(((.((((((......))))))...))))))).....))))))))...(((((.((.....)).))))) ( -40.00) >DroEre_CAF1 4067 120 - 1 GCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGU ..((((((((.(((.....))))))))))).((((((((.......((((.(((.((((((......))))))...))))))).....))))))))...(((((.((.....)).))))) ( -40.00) >consensus GCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUUUCAUUCAUGGCCGGGGCAUACGUAAAUGGAAAAUAUAUGU ..((((((((.(((.....))))))))))).((((((((.......((((.(((.((((((......))))))...))))))).....))))))))...(((((.((.....)).))))) (-40.00 = -40.00 + 0.00)

| Location | 2,871,968 – 2,872,088 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.31 |
| Mean single sequence MFE | -43.49 |
| Consensus MFE | -35.28 |
| Energy contribution | -35.77 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.798490 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 2871968 120 - 27905053 UUGGAGCGCCAACCCCGGGCCCCACCCACAAGUUGGCCAUGCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUU ....((((((......(((.....))).(((...((((..(.((((((((.(((.....))))))))))).)...))))......))))))))).((((((......))))))....... ( -44.80) >DroSec_CAF1 3162 119 - 1 UUGGAGCACCA-CUCCGCGACCCACCCACAAGGUGGCCAUGCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUU ..((((.....-))))(((.((((((.....)))((((..(.((((((((.(((.....))))))))))).)...))))........))))))..((((((......))))))....... ( -44.60) >DroSim_CAF1 3537 119 - 1 UUGGAGCACCA-CUCCGCGACCCACCCACAAGUUGGCCAUGCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUU ..((((.....-))))(((.((((..........((((..(.((((((((.(((.....))))))))))).)...)))).......)))))))..((((((......))))))....... ( -42.43) >DroEre_CAF1 4107 114 - 1 UCGGAGCGCCC------AGCCCCACCCACCAUUUGGCCAUGCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUU (((((.((((.------(((((((..........((((..(.((((((((.(((.....))))))))))).)...)))).......)))).)))))))....)))))............. ( -42.13) >consensus UUGGAGCACCA_CUCCGCGACCCACCCACAAGUUGGCCAUGCGUGCGUGUGUUGGCCAACAAACGCGCACUCUCCGGCCUCAUUAUUGGGCGCUGGCGAUUAUUUGAAAUUGCAAAAGUU ....((((((........................((((..(.((((((((.(((.....))))))))))).)...)))).((....)))))))).((((((......))))))....... (-35.28 = -35.77 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:21 2006