
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,123,142 – 22,123,266 |
| Length | 124 |
| Max. P | 0.936876 |

| Location | 22,123,142 – 22,123,234 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 99.28 |
| Mean single sequence MFE | -16.47 |
| Consensus MFE | -16.47 |
| Energy contribution | -16.47 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.677452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22123142 92 - 27905053 ACAUCCACAUCCGCAUCCGCAUCCUUCUGCCCCAGCUUCUCCUUCGCCUCAUCCUGCACAUCCACUCAGCUCGGCGGUUGCCUUUAAUGCGA ...........(((((..(((.....(((((..((((........((........))..........)))).))))).))).....))))). ( -16.47) >DroSec_CAF1 53259 92 - 1 ACAUCCACAUCCGCAUCCGCAUCCUUCUGCCCCAGCUCCUCCUUCGCCUCAUCCUGCACAUCCACUCAGCUCGGCGGUUGCCUUUAAUGCGA ...........(((((..(((.....(((((..((((........((........))..........)))).))))).))).....))))). ( -16.47) >DroSim_CAF1 45851 92 - 1 ACAUCCACAUCCGCAUCCGCAUCCUUCUGCCCCAGCUCCUCCUUCGCCUCAUCCUGCACAUCCACUCAGCUCGGCGGUUGCCUUUAAUGCGA ...........(((((..(((.....(((((..((((........((........))..........)))).))))).))).....))))). ( -16.47) >consensus ACAUCCACAUCCGCAUCCGCAUCCUUCUGCCCCAGCUCCUCCUUCGCCUCAUCCUGCACAUCCACUCAGCUCGGCGGUUGCCUUUAAUGCGA ...........(((((..(((.....(((((..((((........((........))..........)))).))))).))).....))))). (-16.47 = -16.47 + 0.00)
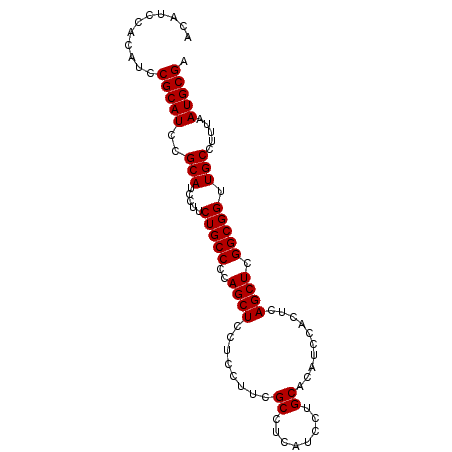



| Location | 22,123,174 – 22,123,266 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 87.75 |
| Mean single sequence MFE | -34.12 |
| Consensus MFE | -24.81 |
| Energy contribution | -24.88 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.627503 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22123174 92 + 27905053 ----UGUG------------CAGGAUGAGGCGAAGG---AGAAGCUGG-GGCAGAAGGAUGCGGAUGCGGAUGUGGAUGUCCUAGGACGGCAACUUGUCCUGUGGCUCCUGC ----...(------------(((((...(.((.(((---(.((.((((-((((...(.((.((....)).)).)...))))))))...(....))).)))).)).))))))) ( -32.70) >DroSec_CAF1 53291 92 + 1 ----UGUG------------CAGGAUGAGGCGAAGG---AGGAGCUGG-GGCAGAAGGAUGCGGAUGCGGAUGUGGAUGUCCUAGGACGGCAACUUGUCCUGUGGCUCCUGC ----...(------------(((((...(.((.(((---(.((.((((-((((...(.((.((....)).)).)...))))))))...(....))).)))).)).))))))) ( -33.10) >DroSim_CAF1 45883 92 + 1 ----UGUG------------CAGGAUGAGGCGAAGG---AGGAGCUGG-GGCAGAAGGAUGCGGAUGCGGAUGUGGAUGUCCUAGGACGGCAACUUGUCCUGUGGCUCCUGC ----...(------------(((((...(.((.(((---(.((.((((-((((...(.((.((....)).)).)...))))))))...(....))).)))).)).))))))) ( -33.10) >DroYak_CAF1 46028 112 + 1 UGGAUGUGGAAGUGGAAAUGCAGGAUGAGGCUAAGGAGGAGGAGCUGGAGGCAGAAGGAUGCGAGUGCGGAUGCGGAUGUCCUAGGACGGCAACUUGUCCUGUGGCUCCUGC ...................((((((....((((((((.(((..((((.(((((...(.((.((....)).)).)...)).)))....))))..))).)))).)))))))))) ( -37.60) >consensus ____UGUG____________CAGGAUGAGGCGAAGG___AGGAGCUGG_GGCAGAAGGAUGCGGAUGCGGAUGUGGAUGUCCUAGGACGGCAACUUGUCCUGUGGCUCCUGC .......................................(((((((((.((((...(.((.((....)).)).)...))))))(((((((....)))))))..))))))).. (-24.81 = -24.88 + 0.06)



| Location | 22,123,174 – 22,123,266 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 87.75 |
| Mean single sequence MFE | -26.18 |
| Consensus MFE | -18.80 |
| Energy contribution | -18.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.936876 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22123174 92 - 27905053 GCAGGAGCCACAGGACAAGUUGCCGUCCUAGGACAUCCACAUCCGCAUCCGCAUCCUUCUGCC-CCAGCUUCU---CCUUCGCCUCAUCCUG------------CACA---- ((((((((...((((.((((((..(((....)))................(((......))).-.)))))).)---)))..))....)))))------------)...---- ( -27.10) >DroSec_CAF1 53291 92 - 1 GCAGGAGCCACAGGACAAGUUGCCGUCCUAGGACAUCCACAUCCGCAUCCGCAUCCUUCUGCC-CCAGCUCCU---CCUUCGCCUCAUCCUG------------CACA---- ((((((((...((((..(((((..(((....)))................(((......))).-.)))))..)---)))..))....)))))------------)...---- ( -25.90) >DroSim_CAF1 45883 92 - 1 GCAGGAGCCACAGGACAAGUUGCCGUCCUAGGACAUCCACAUCCGCAUCCGCAUCCUUCUGCC-CCAGCUCCU---CCUUCGCCUCAUCCUG------------CACA---- ((((((((...((((..(((((..(((....)))................(((......))).-.)))))..)---)))..))....)))))------------)...---- ( -25.90) >DroYak_CAF1 46028 112 - 1 GCAGGAGCCACAGGACAAGUUGCCGUCCUAGGACAUCCGCAUCCGCACUCGCAUCCUUCUGCCUCCAGCUCCUCCUCCUUAGCCUCAUCCUGCAUUUCCACUUCCACAUCCA ((((((((...((((..(((((..(((....)))....((....))....(((......)))...)))))..)))).....))....))))))................... ( -25.80) >consensus GCAGGAGCCACAGGACAAGUUGCCGUCCUAGGACAUCCACAUCCGCAUCCGCAUCCUUCUGCC_CCAGCUCCU___CCUUCGCCUCAUCCUG____________CACA____ (((((((....(((((........))))).(((........)))((....))...))))))).................................................. (-18.80 = -18.80 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:07:16 2006