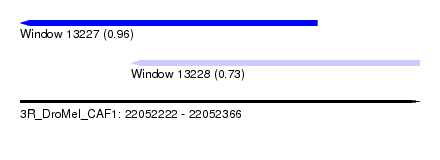
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,052,222 – 22,052,366 |
| Length | 144 |
| Max. P | 0.961614 |

| Location | 22,052,222 – 22,052,329 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.93 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -17.19 |
| Energy contribution | -18.62 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.43 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961614 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
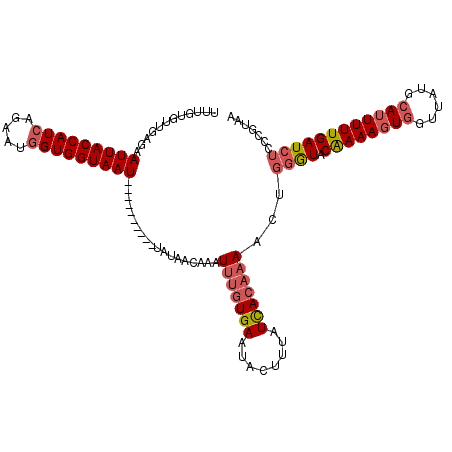
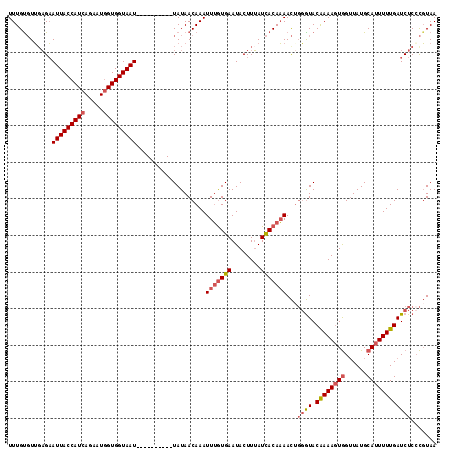
>3R_DroMel_CAF1 22052222 107 - 27905053 UUUGUGUUGAGAAUUACCAUCACCAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAACUGGGUACAAAAGUGGUUAUGCAUUUUUGAUCUCCCGUAA ....(((((..((((((((((.....)))))))))----------).)))))..(((((((........)))))))...(((..((((((((......))))))))....))).... ( -31.30) >DroSec_CAF1 44343 107 - 1 UUUGUGUUCAGAAUUACCAUCAGAAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAACUGGGUACAAAAGUGGUUAUGCAUUUUUGAUCUCACGUAA ((((((..(((((((((((((.....)))))))))----------)........(((((((........))))))).)))..)))))).(.(((((........))))).)...... ( -32.00) >DroSim_CAF1 36983 107 - 1 UUUGCGUUGAGAAUUACCAUCAGAAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAACAGGGUACAAAAGUGGUUAUGCAUUUUUGAUCUCCCGUAA .((((((((..((((((((((.....)))))))))----------).))))...(((((((........)))))))...(((..((((((((......))))))))....))))))) ( -32.00) >DroYak_CAF1 46441 116 - 1 UUUGUCUGGUGUAUUACCAUCAGAAUAGUGGUAAUUAAUGGCAAUUAUAACAAAUUGAUGAAUACUUAAUUACACAAUU-UAUACGAAACUAGGUAUGCAUUUUUGAUCUCCUCUAA ....(((((((......))))))).(((.((..(((((..((.........((((((.(((((.....))).)))))))-)((((........))))))....)))))..)).))). ( -20.90) >consensus UUUGUGUUGAGAAUUACCAUCAGAAUGGUGGUAAU__________UAUAACAAAUUUGUGAAUACUUUAUCACAAAACUGGGUACAAAAGUGGUUAUGCAUUUUUGAUCUCCCGUAA ............(((((((((.....)))))))))...................(((((((........)))))))...((((.((((((((......))))))))))))....... (-17.19 = -18.62 + 1.44)

| Location | 22,052,262 – 22,052,366 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 85.71 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -14.84 |
| Energy contribution | -15.84 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.725905 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
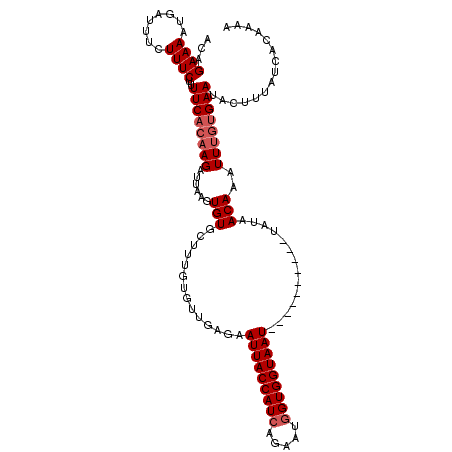
>3R_DroMel_CAF1 22052262 104 - 27905053 ---GAAAAUGAUUUCUUUCUUUUCACAAGAUUAAGUGUGCUUUGUGUUGAGAAUUACCAUCACCAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAA ---(((((.((......)))))))....(((.((((((((....(((((..((((((((((.....)))))))))----------).))))).....))..)))))).)))...... ( -25.30) >DroSec_CAF1 44383 107 - 1 ACAGAAAUUGAUUUAUUUCUUUUCACAAGAUUAAGUGUGCUUUGUGUUCAGAAUUACCAUCAGAAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAA ..((((((......))))))........(((.((((((((....((((...((((((((((.....)))))))))----------)..)))).....))..)))))).)))...... ( -24.30) >DroSim_CAF1 37023 107 - 1 ACAGAAAAUGAUUUCUUUCUUUUCACAAGAUUAAGUGUGCUUUGCGUUGAGAAUUACCAUCAGAAUGGUGGUAAU----------UAUAACAAAUUUGUGAAUACUUUAUCACAAAA ...(((((.((......)))))))....(((.((((((((...))((((..((((((((((.....)))))))))----------).))))..........)))))).)))...... ( -23.80) >DroYak_CAF1 46480 117 - 1 ACAGAAAAUGAUUUCUUUCACUUCUCAAGAUUAAGUGUGCUUUGUCUGGUGUAUUACCAUCAGAAUAGUGGUAAUUAAUGGCAAUUAUAACAAAUUGAUGAAUACUUAAUUACACAA ..((((..(((......))).))))...(((((((((((((...(((((((......)))))))..)))............(((((......)))))....))))))))))...... ( -22.00) >consensus ACAGAAAAUGAUUUCUUUCUUUUCACAAGAUUAAGUGUGCUUUGUGUUGAGAAUUACCAUCAGAAUGGUGGUAAU__________UAUAACAAAUUUGUGAAUACUUUAUCACAAAA ...((((........))))..((((((((......(((..............(((((((((.....)))))))))..............)))..))))))))............... (-14.84 = -15.84 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:05:37 2006