
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,024,504 – 22,024,658 |
| Length | 154 |
| Max. P | 0.924380 |

| Location | 22,024,504 – 22,024,618 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -29.88 |
| Consensus MFE | -26.42 |
| Energy contribution | -26.66 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.609161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22024504 114 + 27905053 UCAUCAGACAUGAAUAAUGCCGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCA ......((....(((((((.((((......)))).).((...((((.(((.....((((((((..........))))))))......)))))))....)).))))))....)). ( -29.20) >DroSec_CAF1 22946 114 + 1 UCAUCAGACAUGAAUAAUGCCGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCA ......((....(((((((.((((......)))).).((...((((.(((.....((((((((..........))))))))......)))))))....)).))))))....)). ( -29.20) >DroSim_CAF1 15009 114 + 1 UCAUCAGACAUGAAUAAUGCCGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAGCAAAUUGCCAACUACCAGUUAUUAAACUCA ......((....(((((((.((((......)))).).((...((((...((....((((((((..........))))))))...))....))))....)).))))))....)). ( -29.40) >DroEre_CAF1 16775 114 + 1 UCAUCGGACAUGAAUAAUGCCGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGCCACAAAUUGCCAGCUACCAGUUAUUAAACUCU .....((.(((.....)))))(((...((((..((((.....))))........(((((((((..........)))))))))))))....)))..................... ( -33.90) >DroYak_CAF1 24241 114 + 1 UCAUCGGACAUGAAUAAUGCUGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCAAGCGCAACAAAUUGCCAACUAGCAGUUAUUAAACUCA ......((....((((((((((((.(((.(((.((((.....))))........(((((.(((..........))).))))).))).))))))....)))).)))))....)). ( -27.70) >consensus UCAUCAGACAUGAAUAAUGCCGGCUAAUGUGUCGCCAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCA ......((....(((((((.((((......)))).).((...((((.(((.....((((((((..........))))))))......)))))))....)).))))))....)). (-26.42 = -26.66 + 0.24)

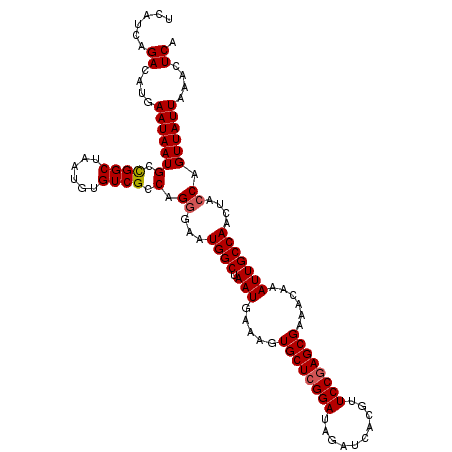

| Location | 22,024,504 – 22,024,618 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -31.44 |
| Consensus MFE | -30.50 |
| Energy contribution | -30.42 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924380 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22024504 114 - 27905053 UGAGUUUAAUAACUGGUAGUUGGCAAUUUGUUUCGCUCGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCGGCAUUAUUCAUGUCUGAUGA ..((((....))))....((((((....(((.(((((((((..........)))))))..........((((.....)))))).)))...)))))).....((((.....)))) ( -31.30) >DroSec_CAF1 22946 114 - 1 UGAGUUUAAUAACUGGUAGUUGGCAAUUUGUUUCGCUCGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCGGCAUUAUUCAUGUCUGAUGA ..((((....))))....((((((....(((.(((((((((..........)))))))..........((((.....)))))).)))...)))))).....((((.....)))) ( -31.30) >DroSim_CAF1 15009 114 - 1 UGAGUUUAAUAACUGGUAGUUGGCAAUUUGCUUCGCUCGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCGGCAUUAUUCAUGUCUGAUGA ..((((....))))....((((((....((..(((((((((..........)))))))..........((((.....))))))..))...)))))).....((((.....)))) ( -29.20) >DroEre_CAF1 16775 114 - 1 AGAGUUUAAUAACUGGUAGCUGGCAAUUUGUGGCGCUCGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCGGCAUUAUUCAUGUCCGAUGA ..((((....))))....((((((....((((..(((((((..........)))))))..........((((.....))))..))))...)))))).....((((.....)))) ( -35.00) >DroYak_CAF1 24241 114 - 1 UGAGUUUAAUAACUGCUAGUUGGCAAUUUGUUGCGCUUGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCAGCAUUAUUCAUGUCCGAUGA ..((((....))))....(((((((((.((((..(((((((..........)))))))..........((((.....)))))))).))).)))))).....((((.....)))) ( -30.40) >consensus UGAGUUUAAUAACUGGUAGUUGGCAAUUUGUUUCGCUCGGAACGUGAUCUAUCCGAGCACUUUCAUUAGCCAUUCCCUGGCGACACAUUAGCCGGCAUUAUUCAUGUCUGAUGA ..((((....))))....(((((((((.((((..(((((((..........)))))))..........((((.....)))))))).))).)))))).....((((.....)))) (-30.50 = -30.42 + -0.08)


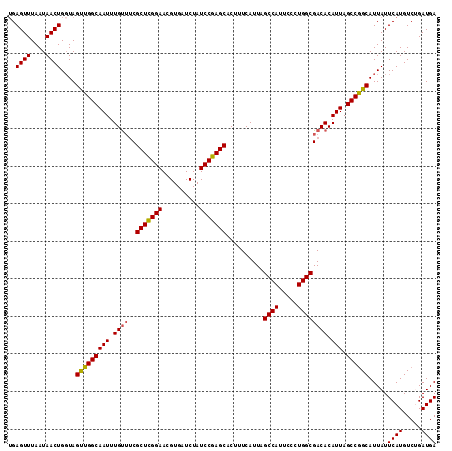
| Location | 22,024,539 – 22,024,658 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 96.30 |
| Mean single sequence MFE | -24.40 |
| Consensus MFE | -20.52 |
| Energy contribution | -20.60 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.550398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22024539 119 + 27905053 CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCAGUCAUUAAUUAUAUUUUACUACCAACAUUUUCGUUAAUAU ..((...((((.(((.....((((((((..........))))))))......)))))))(((...((((....)))).)))...................))................. ( -22.00) >DroSec_CAF1 22981 119 + 1 CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCAGUCAUUAAUUAUAUUUUACUACCAACAUUUUCGUUAAUAU ..((...((((.(((.....((((((((..........))))))))......)))))))(((...((((....)))).)))...................))................. ( -22.00) >DroSim_CAF1 15044 119 + 1 CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAGCAAAUUGCCAACUACCAGUUAUUAAACUCAGUCAUUAAUUAUAUUUUACUACCAACAUUUUCGUUAAUAU ..(..(((((((.((....))(((((((..........)))))))..))).(((((...(((...((((....)))).)))...)))))...............))))..)........ ( -22.50) >DroEre_CAF1 16810 119 + 1 CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGCCACAAAUUGCCAGCUACCAGUUAUUAAACUCUGCCAUUAAUUAUAUUUUAUUACCAACAUUUUCGUUAAUAU ..(..((((..(((((((((((((((((..........)))))))))....(((((...((....((((....))))..))...)))))...))))))))....))))..)........ ( -28.00) >DroYak_CAF1 24276 119 + 1 CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCAAGCGCAACAAAUUGCCAACUAGCAGUUAUUAAACUCAGUCAUUAAUUAUAUUUCAUUACCAACAUUUUCGUUAAUAU ..(..((((..(((((((((((((.(((..........))).)))))....((((((......)))))).......................))))))))....))))..)........ ( -27.50) >consensus CAGGGAAUGGCUAAUGAAAGUGCUCGGAUAGAUCACGUUCCGAGCGAAACAAAUUGCCAACUACCAGUUAUUAAACUCAGUCAUUAAUUAUAUUUUACUACCAACAUUUUCGUUAAUAU ..((...((((.(((.....((((((((..........))))))))......)))))))(((...((((....)))).)))...................))................. (-20.52 = -20.60 + 0.08)

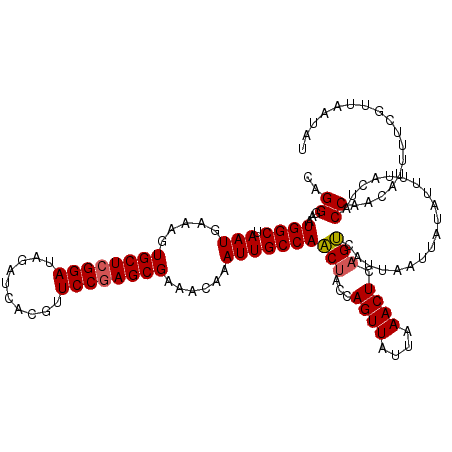

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:05:04 2006