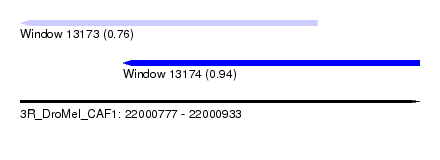
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 22,000,777 – 22,000,933 |
| Length | 156 |
| Max. P | 0.935957 |

| Location | 22,000,777 – 22,000,893 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.66 |
| Mean single sequence MFE | -42.22 |
| Consensus MFE | -33.43 |
| Energy contribution | -33.99 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.757671 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22000777 116 - 27905053 AAUCGCCAGGUGAACAGGCC---GGA-GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGACAAGCCUGGGCACUUAUUUGCCGAACGCUUCUGCUUGGGCA .......(((((..(((((.---..(-((....)))..(((((((.(((((((.............))).))))))))))).)))))..)))))...((((.((.((....)))).)))) ( -41.22) >DroSim_CAF1 37155 119 - 1 AAUCGCCAGGUGAACAGGCCAGUGGA-GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGACGAGCCUGGGCACUUAUUUGCCGAACGCUUCUGCCUGGGCA ..(((((((((((((((((.((((..-.)))).))..)))))))))(((((((.............))).)))).))).)))(((..((((.......((.....))...))))..))). ( -44.92) >DroEre_CAF1 42809 120 - 1 AAUAGCAGGGUGAACAGGCCAGUGAAGGCACUUUCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGAAGAGCCUAGGCACUUAUUUGCCGAACGCUUCUGCCUGUGCA ....(((((.(((((((...((((....)))).....)))))))))))).........((((((..(.(((((((...(((.((....)).)))..))))))).)((....)).)))))) ( -41.70) >DroYak_CAF1 41436 120 - 1 AAUCGCCAGGUGAACAGGCCAGUGAAGGCACUUUCUGCUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUGGCCAUUGGACGAGCAUGGGCACUUAUUUGCCGAACGCUUCUGCCUGGGCA ....((((((((((((((((......))).(.....)))))))))))((((((.............))))))....((..((((.(.((((......)))).)..))))...))..))). ( -41.02) >consensus AAUCGCCAGGUGAACAGGCCAGUGAA_GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGACGAGCCUGGGCACUUAUUUGCCGAACGCUUCUGCCUGGGCA ....(((((((((((((((.((((....)))).))..))))))))))((((((.............))).)))...((..((((...((((......))))....))))...))..))). (-33.43 = -33.99 + 0.56)



| Location | 22,000,817 – 22,000,933 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.13 |
| Mean single sequence MFE | -36.73 |
| Consensus MFE | -29.45 |
| Energy contribution | -31.07 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935957 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 22000817 116 - 27905053 AAAAACAAACAUAGCUAGUGAGGAGCCUAAAAGCUUCUUAAAUCGCCAGGUGAACAGGCC---GGA-GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGAC .........(((.(((((((((((((......))))))).....(.(((((((((((...---..(-((....))).))))))))))).)................))).))).)))... ( -35.10) >DroSim_CAF1 37195 119 - 1 AAAAACAAACAUAGCUAGUGAGGAGCCUAAAAGCUUCUUAAAUCGCCAGGUGAACAGGCCAGUGGA-GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGAC .........(((.(((((((((((((......))))))).....(.(((((((((((((.((((..-.)))).))..))))))))))).)................))).))).)))... ( -36.10) >DroEre_CAF1 42849 120 - 1 UUUAACAAACAUGGCUAGUGAGGAGUCCAUAAGCUUCUUAAAUAGCAGGGUGAACAGGCCAGUGAAGGCACUUUCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGAA .........(((.((((.((((((((......))))))))..)))).((((((((((...((((....)))).....))))))))))((((((.............))).))).)))... ( -34.92) >DroYak_CAF1 41476 120 - 1 UUUAACAAACAUGGCUAGUGAGGAGCCUAUAAGCUUCUUAAAUCGCCAGGUGAACAGGCCAGUGAAGGCACUUUCUGCUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUGGCCAUUGGAC .........(((((((((((((((((......))))))).....(.((((((((((((((......))).(.....)))))))))))).)................)))))))).))... ( -40.80) >consensus AAAAACAAACAUAGCUAGUGAGGAGCCUAAAAGCUUCUUAAAUCGCCAGGUGAACAGGCCAGUGAA_GCACUUGCUACUGUUUACCUGCCAGUUAAUUUGCAUAAUGCUCGGCAGUGGAC .........(((.(((((((((((((......))))))).....(.(((((((((((((.((((....)))).))..))))))))))).)................))).))).)))... (-29.45 = -31.07 + 1.62)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:04:46 2006