
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 21,641,498 – 21,641,658 |
| Length | 160 |
| Max. P | 0.822133 |

| Location | 21,641,498 – 21,641,618 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.00 |
| Mean single sequence MFE | -59.28 |
| Consensus MFE | -49.48 |
| Energy contribution | -49.82 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.611662 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21641498 120 + 27905053 CAGGAGGCCAAGAAGGCGUCGGACACUCAGGCACCAGCUGCUCUGGCUGCUGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAAGCCAUUGCUGCGGCCAAGGCCGCUGAGGCGGCGCUG (((..((((.....((((..((...(((.((((.(((((.....))))).)))).(((((.((......)).))))).)))....))..))))..))))...(((((....))))).))) ( -54.00) >DroSec_CAF1 130844 120 + 1 CAGGAGGCCAAGAAGGCGUCGGACACUCAGGCACCUGCGGCUCUGGCUGCUGCCCGCCAGGUGAAACACCACCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCCGCUGAGGCGGCGCUG (((..((((....(((.(((.(.....).))).)))(((((..(((((....((.((((((((......)))))))).).)..)))))..)))))))))...(((((....))))).))) ( -61.40) >DroSim_CAF1 127445 120 + 1 CAGGAGGCCAAGAAGGCGUCGGACACUCAGGCACCUGCGGCUCUGGCUGCUGCCCGCCAGGUGAAACACCACCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCCGCCGAGGCGGCGCUG (((..((((....(((.(((.(.....).))).)))(((((..(((((....((.((((((((......)))))))).).)..)))))..)))))))))...(((((....))))).))) ( -62.50) >DroEre_CAF1 127564 120 + 1 CAGGAGGCCAAGAAGGCGUCGGACACCCAGGCACCUGCGGCCCUGGCUGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAAGCCGCCGAGGCGGCGCUG (((..((((....(((.(((((....)).))).)))(((((..(((((..(..(.(((((.((......)).))))).).)..)))))..)))))))))...(((((....))))).))) ( -59.60) >DroYak_CAF1 128095 120 + 1 CAGGAGGCCAAGAAGGCGUCGGACACCCAGGCACCUGCGGCUCUGGCCGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCAGCCGAGGCGGCGCUG .....((((....(((.(((((....)).))).)))(((((..(((((..(..(.(((((.((......)).))))).).)..)))))..)))))))))..(((.((((...))))))). ( -57.00) >DroAna_CAF1 110454 120 + 1 AAGGAGGCCAAGCUGGCCUCGGAGUCCCAGCAGCCGGCUGCAUUGGCCGCCGCCUACCAGGUGCGCGUCCAGCUGGCCGACAAGGCUGUCCAAGCGGCCAAGGCCGCAGAGGCAGCUCUG ..(..(((((.(((((((..((....)).((((....))))...)))(((((((.....)))).)))..)))))))))..).((((((((...(((((....)))))...)))))).)). ( -61.20) >consensus CAGGAGGCCAAGAAGGCGUCGGACACCCAGGCACCUGCGGCUCUGGCUGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCCGCCGAGGCGGCGCUG (((..((((....(((.(((((....)).))).)))(((((..(((((.....(.(((((.((......)).))))).)....)))))..)))))))))...(((((....))))).))) (-49.48 = -49.82 + 0.34)



| Location | 21,641,538 – 21,641,658 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.39 |
| Mean single sequence MFE | -54.25 |
| Consensus MFE | -33.60 |
| Energy contribution | -33.30 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.822133 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21641538 120 + 27905053 CUCUGGCUGCUGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAAGCCAUUGCUGCGGCCAAGGCCGCUGAGGCGGCGCUGGCGGGGAAACAACAACUUAUGGAGCAGCUGCAGGACGAGG .(((.((.((((((((((.((((...))))(((((((((....(((....))).))))))..(((((....)))))))))))))(....).............))))).)).)))..... ( -57.40) >DroGri_CAF1 108022 120 + 1 CCGCGGAGGUCGCCCGCAUGGUUAAGAAGCAGCUGGCGGAUCGAGCAUAUGCAGCUGCCAAAGCAGCGGAGGCGGCACUCUCUGGCAAGCAACAGAAUGUGGAGCAGCUAGAGCAGGAGA ((((.((((((((((((((((((.....((.....))......))).))))).(((((....))))).).)))))).)))((((((..((..((.....))..)).)))))))).))... ( -44.90) >DroSim_CAF1 127485 120 + 1 CUCUGGCUGCUGCCCGCCAGGUGAAACACCACCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCCGCCGAGGCGGCGCUGGCGGGAAAACAACAGCUUAUGGAGCAGCUGCAGGACGAGG .(((.((.(((((((((((((((......))))))))......((((........))))...(((((....)))))((((..(......)..))))....)).))))).)).)))..... ( -57.60) >DroEre_CAF1 127604 120 + 1 CCCUGGCUGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAAGCCGCCGAGGCGGCGCUGGCGGGGAAGCAACAGAUUAUGGAGCAGCUGCAGGACGAGG .((((((((((.((((((.((((...))))(((((((((....(((....))).))))))..(((((....)))))))))))))).......((.....))).)))))..))))...... ( -59.60) >DroYak_CAF1 128135 120 + 1 CUCUGGCCGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCAGCCGAGGCGGCGCUGGCGGGGAAACAACAGAUCAUGGAACAGCUGCAGGACGAGG .((((.((((((((((((.((((...))))((((((((.....)))))..))).((((.......)))).))))).)).)))))(....)..))))........................ ( -50.90) >DroAna_CAF1 110494 120 + 1 CAUUGGCCGCCGCCUACCAGGUGCGCGUCCAGCUGGCCGACAAGGCUGUCCAAGCGGCCAAGGCCGCAGAGGCAGCUCUGGCCGGGAAGGAGCAGAUAGUCGAACAGCUGCAGGAAGAGG ..((..((((((((.....)))).))...((((((..((((...((((((...(((((....)))))...))))))((((.((.....))..))))..))))..))))))..))..)).. ( -55.10) >consensus CUCUGGCUGCCGCCCGCCAGGUGAAACACCAGCUGGCCGAGAAGGCCAUUGCUGCGGCCAAGGCCGCCGAGGCGGCGCUGGCGGGGAAACAACAGAUUAUGGAGCAGCUGCAGGACGAGG .(((.(((((((((((((((.((......)).)))))................(((((....))))).).))))))...))).))).................................. (-33.60 = -33.30 + -0.30)



| Location | 21,641,538 – 21,641,658 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.39 |
| Mean single sequence MFE | -51.47 |
| Consensus MFE | -35.09 |
| Energy contribution | -36.82 |
| Covariance contribution | 1.73 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.794716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
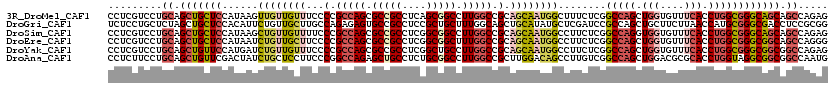
>3R_DroMel_CAF1 21641538 120 - 27905053 CCUCGUCCUGCAGCUGCUCCAUAAGUUGUUGUUUCCCCGCCAGCGCCGCCUCAGCGGCCUUGGCCGCAGCAAUGGCUUUCUCGGCCAGCUGGUGUUUCACCUGGCGGGCAGCAGCCAGAG .(((.....((.(((((.((..((((..(((((.(...(((((.(((((....))))).))))).).)))))..)))).....(((((.(((....))).)))))))))))).))..))) ( -52.60) >DroGri_CAF1 108022 120 - 1 UCUCCUGCUCUAGCUGCUCCACAUUCUGUUGCUUGCCAGAGAGUGCCGCCUCCGCUGCUUUGGCAGCUGCAUAUGCUCGAUCCGCCAGCUGCUUCUUAACCAUGCGGGCGACCUCCGCGG ....(((((((.((.((..((.....))..))..)).)))(((.(.((((..(((((.((.((((((((.....((.......))))))))))....)).)).))))))).)))).)))) ( -35.90) >DroSim_CAF1 127485 120 - 1 CCUCGUCCUGCAGCUGCUCCAUAAGCUGUUGUUUUCCCGCCAGCGCCGCCUCGGCGGCCUUGGCCGCAGCAAUGGCCUUCUCGGCCAGGUGGUGUUUCACCUGGCGGGCAGCAGCCAGAG .(((.....((.(((((.((....(((((((((...(.(((((.(((((....))))).))))).).))))))))).......(((((((((....)))))))))))))))).))..))) ( -60.90) >DroEre_CAF1 127604 120 - 1 CCUCGUCCUGCAGCUGCUCCAUAAUCUGUUGCUUCCCCGCCAGCGCCGCCUCGGCGGCUUUGGCCGCAGCAAUGGCCUUCUCGGCCAGCUGGUGUUUCACCUGGCGGGCGGCAGCCAGGG ......(((((.(((((.((......((((((......(((((.(((((....))))).))))).)))))).(((((.....)))))(((((((...)))).)))))))))).).)))). ( -57.40) >DroYak_CAF1 128135 120 - 1 CCUCGUCCUGCAGCUGUUCCAUGAUCUGUUGUUUCCCCGCCAGCGCCGCCUCGGCUGCCUUGGCCGCAGCAAUGGCCUUCUCGGCCAGCUGGUGUUUCACCUGGCGGGCGGCGGCCAGAG .(((.....(((((...((...))...)))))....(((((.((.(((((.(((((.....)))))((((..(((((.....)))))))))...........))))))))))))...))) ( -49.10) >DroAna_CAF1 110494 120 - 1 CCUCUUCCUGCAGCUGUUCGACUAUCUGCUCCUUCCCGGCCAGAGCUGCCUCUGCGGCCUUGGCCGCUUGGACAGCCUUGUCGGCCAGCUGGACGCGCACCUGGUAGGCGGCGGCCAAUG ((.((.((((((((((..((((...(((.(((....(((((((.(((((....))))).)))))))...))))))....))))..)))))((.......))..))))).)).))...... ( -52.90) >consensus CCUCGUCCUGCAGCUGCUCCAUAAUCUGUUGCUUCCCCGCCAGCGCCGCCUCGGCGGCCUUGGCCGCAGCAAUGGCCUUCUCGGCCAGCUGGUGUUUCACCUGGCGGGCGGCAGCCAGAG .........((.(((((((......((((((((...(.(((((.(((((....))))).))))).).))))))))........(((((.(((....))).)))))))))))).))..... (-35.09 = -36.82 + 1.73)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:38 2006