
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 21,259,586 – 21,259,843 |
| Length | 257 |
| Max. P | 0.952423 |

| Location | 21,259,586 – 21,259,692 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.11 |
| Mean single sequence MFE | -22.45 |
| Consensus MFE | -13.00 |
| Energy contribution | -12.81 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.566798 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21259586 106 - 27905053 AAGGUAUCCAAGCUUUAGAUAGCUUGAAUAUUAUUUGG-GAAAAAGUUCACGAGUUGCUUUUUAGCAAAUGAUUCUAUUCUAGAAGGUGAUUUAAGUAAAGAUACAU------------- ...(((((((((((......))))))....((((((..-(((....)))..((((..((((((((..((((....))))))))))))..)))))))))).)))))..------------- ( -24.90) >DroSec_CAF1 84454 94 - 1 AAGGUAUCCAAACU----------UAUAUUUUAUUUGG-GAUAAAGUUCACGAGUUGCUGUUUAGAUGUUGAUUU--UUCUAGAAGGUGACUUAAGUAAAGAUACAU------------- ...(((((...(((----------(..((((((((...-))))))))....((((..((.((((((.........--.)))))).))..))))))))...)))))..------------- ( -20.80) >DroSim_CAF1 87178 94 - 1 AAGGUAUCCAAACU----------UAUAUUUUAUAUGG-GAUAAAGGUCACGAGUUGCUGUUUAGCUAUUGAUUU--UUCUAGAAGGUGACUUAAGUAAAGAUACAU------------- ...(((((...(((----------(..........(((-((...((((((..(((((.....)))))..))))))--))))).(((....)))))))...)))))..------------- ( -18.10) >DroYak_CAF1 93975 118 - 1 AAUGUAUCCACAAUUUAAAUAACUUGUGAUUUAUUUGGGGAUAACGUCCGUGAGUGGCUUUUUAGUUAUUGAUAU--UUUUAGAAGGUGACUUAAGUAAAGUUGCAUUUAUUUAAACGAC .............(((((((((..((..((((......((((...)))).(((((.(((((((((..((....))--..))))))))).)))))....))))..)).))))))))).... ( -26.00) >consensus AAGGUAUCCAAACU__________UAUAUUUUAUUUGG_GAUAAAGUUCACGAGUUGCUGUUUAGCUAUUGAUUU__UUCUAGAAGGUGACUUAAGUAAAGAUACAU_____________ ...(((((...........................(((.........))).((((..((((((((..............))))))))..)))).......)))))............... (-13.00 = -12.81 + -0.19)


| Location | 21,259,692 – 21,259,804 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 95.72 |
| Mean single sequence MFE | -28.57 |
| Consensus MFE | -24.73 |
| Energy contribution | -25.23 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.599929 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
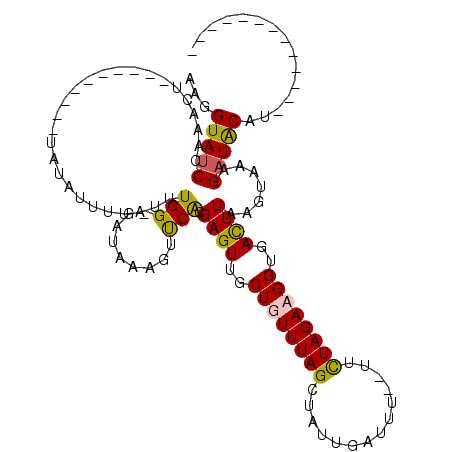
>3R_DroMel_CAF1 21259692 112 - 27905053 GUUUUUUUAUCCGACAUCAUGUGCAUGAUGUUAAUGCGCUAAGUGCAAUGCGAAACAAGGGGA-AAACCCAAAUGAUUUAUGGCAAGCGGAUCACGUGAGCCAAAUUUUU-AUA (((((.......((((((((....))))))))..(((((...)))))....)))))..(((..-...)))..........((((..(((.....)))..)))).......-... ( -30.40) >DroSec_CAF1 84548 113 - 1 GUUU-UUUAUCCGACAUCAUGUGCAUGAUGUUAAUGCGCUAAGUGCAAUGCGAAACAAGGGAACAAACCCAAAUGAUUUAUGGCAAGCGGAUCACGUGAGCCAAAUUUUUGAUA ((((-(.(((..((((((((....))))))))..(((((...)))))))).)))))..(((......)))..........((((..(((.....)))..))))........... ( -29.70) >DroSim_CAF1 87272 112 - 1 GUUU-UUUAUCCGACAUCAUGUGCAUGAUGUUAAUGCGCUAAGUGCAAUGCGAAACAAGGGGA-AAACCCAAAUGAUUUAUGGCAAGCGAAUCACGUGAGCCAAAUUUUUGAUA ((((-(.(((..((((((((....))))))))..(((((...)))))))).)))))..(((..-...)))..........((((..(((.....)))..))))........... ( -29.50) >DroYak_CAF1 94093 111 - 1 GUUU-UUUAUCCGACAUCAUGUGCAUGAUGUUAAUGCGCUAAGUGCAAUGCGAAACAAGUGAA-AAACCCAAAUGAUUUAUGGCAAGGGGGUCACGUAAGCCAAAUUUUU-AUA ((((-(.(((..((((((((....))))))))..(((((...)))))))).)))))..((((.-...(((...((........))..))).))))...............-... ( -24.70) >consensus GUUU_UUUAUCCGACAUCAUGUGCAUGAUGUUAAUGCGCUAAGUGCAAUGCGAAACAAGGGAA_AAACCCAAAUGAUUUAUGGCAAGCGGAUCACGUGAGCCAAAUUUUU_AUA ((((........((((((((....))))))))..(((((...))))).....))))..(((......)))..........((((..(((.....)))..))))........... (-24.73 = -25.23 + 0.50)


| Location | 21,259,731 – 21,259,843 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.20 |
| Mean single sequence MFE | -32.36 |
| Consensus MFE | -10.48 |
| Energy contribution | -14.00 |
| Covariance contribution | 3.52 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.32 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952423 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
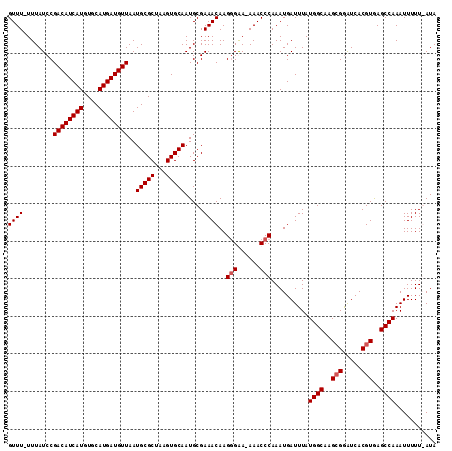
>3R_DroMel_CAF1 21259731 112 - 27905053 CAUCAUUGUUUGGUUACUUCUCUGGGUUU-UUGUGUCCACGUUUUUUUAUCCGACAUCAUGUGCAUG--AU--GUUAAUGCGCUAAGUGCAAUGCGA--AACAAGGGGA-AAACCCAAAU .......((((((....((((((..((((-(.((((................((((((((....)))--))--)))..(((((...)))))))))))--)))..)))))-)...)))))) ( -28.40) >DroSec_CAF1 84588 111 - 1 CAUCAUUGUUGGGUUACUUUUCUGGGUU--UUGCGUCCACGUUU-UUUAUCCGACAUCAUGUGCAUG--AU--GUUAAUGCGCUAAGUGCAAUGCGA--AACAAGGGAACAAACCCAAAU ........(((((((..((..((..(((--((((((........-.......((((((((....)))--))--)))..(((((...)))))))))))--))).))..))..))))))).. ( -36.30) >DroSim_CAF1 87312 110 - 1 CAUCAUUGUUGGGUUACUUUUCUGGGUU--UUGCGUCCACGUUU-UUUAUCCGACAUCAUGUGCAUG--AU--GUUAAUGCGCUAAGUGCAAUGCGA--AACAAGGGGA-AAACCCAAAU ........(((((((..((..((..(((--((((((........-.......((((((((....)))--))--)))..(((((...)))))))))))--)))..))..)-)))))))).. ( -36.80) >DroYak_CAF1 94132 110 - 1 CAUCAUUGUUGAGUUACUUUUCUGGGUU--UUGGGUUCACGUUU-UUUAUCCGACAUCAUGUGCAUG--AU--GUUAAUGCGCUAAGUGCAAUGCGA--AACAAGUGAA-AAACCCAAAU ..(((....)))..........((((((--((....((((((((-(.(((..((((((((....)))--))--)))..(((((...)))))))).))--)))..)))))-)))))))... ( -27.60) >DroAna_CAF1 78420 98 - 1 CAGCUCUGA----GGACUUUUCGGGAUCCGUUGGGU----GUCU-UUAAUG------------CAGGGGAUCCACUAAUGUGCCGAGUGCACUGGGAUAAAUCAGGGCC-AAGUAAGCAU ..(((((((----.....(..((((((((.(((.((----....-...)).------------))).))))))......((((.....))))))..)....))))))).-.......... ( -32.70) >consensus CAUCAUUGUUGGGUUACUUUUCUGGGUU__UUGCGUCCACGUUU_UUUAUCCGACAUCAUGUGCAUG__AU__GUUAAUGCGCUAAGUGCAAUGCGA__AACAAGGGAA_AAACCCAAAU ........(((((((...(((((..(((.(((((((..........................((((...........))))((.....)).))))))).)))..)))))..))))))).. (-10.48 = -14.00 + 3.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:27 2006