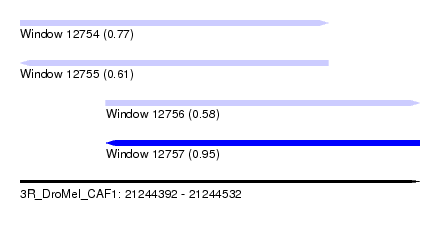

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 21,244,392 – 21,244,532 |
| Length | 140 |
| Max. P | 0.948179 |

| Location | 21,244,392 – 21,244,500 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.40 |
| Mean single sequence MFE | -22.91 |
| Consensus MFE | -15.79 |
| Energy contribution | -15.77 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.771816 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21244392 108 + 27905053 CCCG---------C-UUUUCCUGUCUUUUGAGCCAACUUCAAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU--UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUUGGUUGCC ...(---------(-....((.(((...((((.(((.......)))..))))...((((.((((((((..........--.........)))))))).(.....)))))))).))..)). ( -23.01) >DroSec_CAF1 69155 107 + 1 CCCG---------C-CUUUCCUGUCUUUUAAGCCAACUUCAAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUU---UUUUAUCCUUUGGCGAAAGACCAUCGGUCGAUUGGUUGCC ...(---------(-....((.(((......(((((.(((.....))).....)))))..((((((((.........---.........)))))))).((((...))))))).))..)). ( -21.47) >DroEre_CAF1 73210 108 + 1 CCCG---------C-UUUUCCUGUCUUUUGAGCCAACUUCAAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU--UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUCGGUUGGC ....---------.-................(((((((((.....))......((((((.((((((((..........--.........)))))))).(.....)))))))..))))))) ( -25.21) >DroWil_CAF1 72500 107 + 1 UUCGUUUUUGGUUC-AUUUCC-------UAUGCCAACUUCAAAUUGAACUCAUUUGGCCAGUUGCCAAUUAACUUUUU-----CCCCUUUUAGCCAAAGAGCAAUGGCCGAUUUGUAACC .........(((((-(..((.-------...((((..(((.....)))(((.((((((.(((((.....)))))....-----.........)))))))))...)))).))..)).)))) ( -18.54) >DroYak_CAF1 78575 110 + 1 CCCG---------C-UUUUCCUGUCUUUUGAGCCAACUUCAAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUUUUUUUUAUCCUUUGGCAAAAGACCAUCGGCCGAUUGGUUGCC ...(---------(-....((.(((...((((.(((.......)))..))))...((((.((((((((.....................)))))))).(.....)))))))).))..)). ( -23.20) >DroAna_CAF1 65158 109 + 1 CCAC---------CGUUCUUCUGCCU-UUCAGCCAACUUCAAAUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU-CGCUCAUCCUUUGGCAGAGGAGCAUUGCUCGAUUGGUUACC ..((---------((((((((((((.-...(((....(((.....))).....(((((.....)))))..........-.)))........)))))))))))(((....))).))).... ( -26.00) >consensus CCCG_________C_UUUUCCUGUCUUUUGAGCCAACUUCAAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU__UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUUGGUUGCC ..............................((((((.(((.....))).....((((((.((((((((.....................)))))))).(.....)))))))))))))... (-15.79 = -15.77 + -0.03)


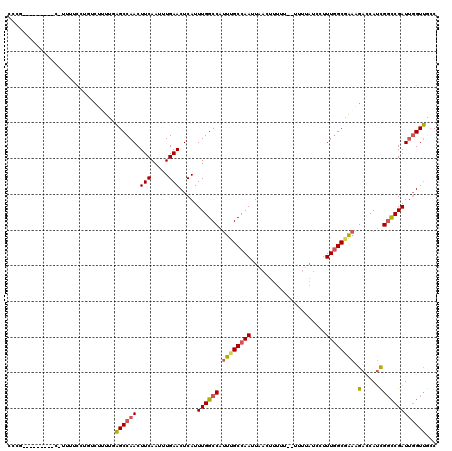
| Location | 21,244,392 – 21,244,500 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.40 |
| Mean single sequence MFE | -26.37 |
| Consensus MFE | -15.95 |
| Energy contribution | -15.87 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.609720 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21244392 108 - 27905053 GGCAACCAAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA--AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUUGAAGUUGGCUCAAAAGACAGGAAAA-G---------CGGG .((..((....))((....((((((.(((((..(((....--.....)))..)))))...(((((((.....(((.....))).))))).)))))))).))....-)---------)... ( -23.70) >DroSec_CAF1 69155 107 - 1 GGCAACCAAUCGACCGAUGGUCUUUCGCCAAAGGAUAAAA---AAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUUGAAGUUGGCUUAAAAGACAGGAAAG-G---------CGGG .....((.(((....))).((((((((((((..(((....---....)))..)))))....((((((.....(((.....))).))))))..........)))))-)---------))). ( -26.40) >DroEre_CAF1 73210 108 - 1 GCCAACCGAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA--AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUUGAAGUUGGCUCAAAAGACAGGAAAA-G---------CGGG .....(((.....((....((((((.(((((..(((....--.....)))..)))))...(((((((.....(((.....))).))))).)))))))).))....-.---------))). ( -27.10) >DroWil_CAF1 72500 107 - 1 GGUUACAAAUCGGCCAUUGCUCUUUGGCUAAAAGGGG-----AAAAAGUUAAUUGGCAACUGGCCAAAUGAGUUCAAUUUGAAGUUGGCAUA-------GGAAAU-GAACCAAAAACGAA ((((.((..((((((((((((..((((((........-----....))))))..))))).)))))....((.(((.....))).))......-------.))..)-)))))......... ( -26.40) >DroYak_CAF1 78575 110 - 1 GGCAACCAAUCGGCCGAUGGUCUUUUGCCAAAGGAUAAAAAAAAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUUGAAGUUGGCUCAAAAGACAGGAAAA-G---------CGGG .((..((..(((((((((((((.((((((((..(((...........)))..)))))))).)))).......(((.....)))))))))).....))..))....-)---------)... ( -26.70) >DroAna_CAF1 65158 109 - 1 GGUAACCAAUCGAGCAAUGCUCCUCUGCCAAAGGAUGAGCG-AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAUUUGAAGUUGGCUGAA-AGGCAGAAGAACG---------GUGG ....(((....((((..(((((.(((......))).)))))-......(((.(((((.....))))).)))))))............(((...-.)))........)---------)).. ( -27.90) >consensus GGCAACCAAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA__AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUUGAAGUUGGCUCAAAAGACAGGAAAA_G_________CGGG .....((....((((...))))....(((((.................(((.(((((.....))))).))).(((.....))).)))))..........))................... (-15.95 = -15.87 + -0.08)

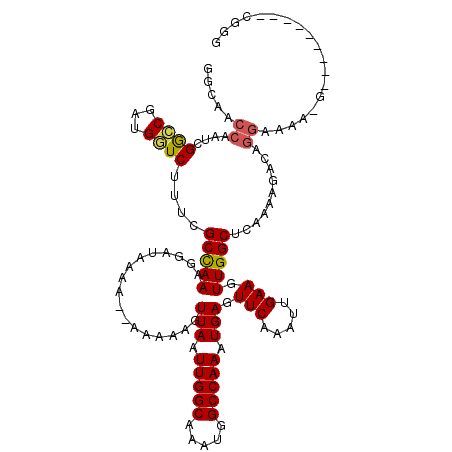

| Location | 21,244,422 – 21,244,532 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.81 |
| Mean single sequence MFE | -26.69 |
| Consensus MFE | -14.83 |
| Energy contribution | -15.42 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.580941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21244422 110 + 27905053 AAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU--UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUUGGUUGCCCCUUUCUGCAGGACCA---GG-CCAACACACCCACC---- .............((((((.((((((((..........--.........))))))))...((....((.((..((.....))..)).)).))....---))-))))..........---- ( -25.01) >DroSec_CAF1 69185 109 + 1 AAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUU---UUUUAUCCUUUGGCGAAAGACCAUCGGUCGAUUGGUUGCCCCUUUCUGCAGGAUCA---GG-CCAACCCACCCACC---- .............((((((.((((((((.........---.........)))))))).((((...))))((((..((((........)))))))).---))-))))..........---- ( -26.37) >DroEre_CAF1 73240 110 + 1 AAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU--UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUCGGUUGGCCCUUUCUGCGGGGCCA---GG-CCAACCCACCCACC---- .............((((((.((((((((..........--.........)))))))).(((((((....))).))))((((((......)))))).---))-))))..........---- ( -35.21) >DroWil_CAF1 72532 104 + 1 AAAUUGAACUCAUUUGGCCAGUUGCCAAUUAACUUUUU-----CCCCUUUUAGCCAAAGAGCAAUGGCCGAUUUGUAACCUC---CCGUGCAGCUA---CU-UCACCUCACCCCUU---- ....((((.....(((((((.(((((............-----...............).))))))))))).(((((.....---...)))))...---.)-)))...........---- ( -14.43) >DroYak_CAF1 78605 113 + 1 AAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUUUUUUUUAUCCUUUGGCAAAAGACCAUCGGCCGAUUGGUUGCCCCUUUCUGCGGGGCCA---GGACCAGCCCACCCACC---- .............((((((.((((((((.....................)))))))).(.....)))))))(((((((((((.......)))))..---.))))))..........---- ( -31.70) >DroAna_CAF1 65188 118 + 1 AAAUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU-CGCUCAUCCUUUGGCAGAGGAGCAUUGCUCGAUUGGUUACCCCUUUUCGCAGGGCAACUGCG-CCCACCCACCCACUGUUC .....((((.....((((((((((((((..........-..........)))))))).((((...))))...))))))............((((......)-)))...........)))) ( -27.45) >consensus AAUUUGAACUCAUUUGGCCAUUUGCCAAUUAACUUUUU__UUUUAUCCUUUGGCGAAAGACCAUCGGCCGAUUGGUUGCCCCUUUCUGCAGGGCCA___GG_CCAACCCACCCACC____ ......(((.((.((((((.((((((((.....................)))))))).(.....))))))).)))))(.((((......)))).)......................... (-14.83 = -15.42 + 0.59)



| Location | 21,244,422 – 21,244,532 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.81 |
| Mean single sequence MFE | -35.72 |
| Consensus MFE | -22.14 |
| Energy contribution | -22.20 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.38 |
| SVM RNA-class probability | 0.948179 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21244422 110 - 27905053 ----GGUGGGUGUGUUGG-CC---UGGUCCUGCAGAAAGGGGCAACCAAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA--AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUU ----(((.(((..((((.-((---(..((.....))..))).))))..))).)))..(((((.((.(((((..(((....--.....)))..))))).)).))))).............. ( -32.90) >DroSec_CAF1 69185 109 - 1 ----GGUGGGUGGGUUGG-CC---UGAUCCUGCAGAAAGGGGCAACCAAUCGACCGAUGGUCUUUCGCCAAAGGAUAAAA---AAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUU ----(((.(((.(((((.-((---(..((.....))..))).))))).))).)))..(((((.((.(((((..(((....---....)))..))))).)).))))).............. ( -36.40) >DroEre_CAF1 73240 110 - 1 ----GGUGGGUGGGUUGG-CC---UGGCCCCGCAGAAAGGGCCAACCGAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA--AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUU ----(((.(((.((((((-((---(.((...)).....))))))))).))).)))..(((((.((.(((((..(((....--.....)))..))))).)).))))).............. ( -41.70) >DroWil_CAF1 72532 104 - 1 ----AAGGGGUGAGGUGA-AG---UAGCUGCACGG---GAGGUUACAAAUCGGCCAUUGCUCUUUGGCUAAAAGGGG-----AAAAAGUUAAUUGGCAACUGGCCAAAUGAGUUCAAUUU ----......((((.(.(-.(---(((((......---..)))))).....((((((((((..((((((........-----....))))))..))))).)))))...).).)))).... ( -27.50) >DroYak_CAF1 78605 113 - 1 ----GGUGGGUGGGCUGGUCC---UGGCCCCGCAGAAAGGGGCAACCAAUCGGCCGAUGGUCUUUUGCCAAAGGAUAAAAAAAAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUU ----.((.(((.(..((((..---..(((((.......))))).))))..).))).))((((.((((((((..(((...........)))..)))))))).))))............... ( -41.30) >DroAna_CAF1 65188 118 - 1 GAACAGUGGGUGGGUGGG-CGCAGUUGCCCUGCGAAAAGGGGUAACCAAUCGAGCAAUGCUCCUCUGCCAAAGGAUGAGCG-AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAUUU ((((...(((..(.((..-..)).)..)))(((......((....))......))).(((((.(((......))).)))))-......(((.(((((.....))))).)))))))..... ( -34.50) >consensus ____GGUGGGUGGGUUGG_CC___UGGCCCCGCAGAAAGGGGCAACCAAUCGGCCGAUGGUCUUUCGCCAAAGGAUAAAA__AAAAAGUUAAUUGGCAAAUGGCCAAAUGAGUUCAAAUU .....((.(((.((((((........(((((.......)))))..)))))).))).)).((((((.....)))))).........((.(((.(((((.....))))).))).))...... (-22.14 = -22.20 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:05 2006