
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 21,214,165 – 21,214,305 |
| Length | 140 |
| Max. P | 0.962425 |

| Location | 21,214,165 – 21,214,279 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 79.15 |
| Mean single sequence MFE | -42.17 |
| Consensus MFE | -23.90 |
| Energy contribution | -25.93 |
| Covariance contribution | 2.03 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.77 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962425 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21214165 114 + 27905053 ACUUUGAAGCCGCUGCCGAUGGUGAAGCUAUAGCAAAAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGGAUCGUUCGACGGAAGUGGU ........((((((.((((((((...((((((((....))))))))((.(((...(((....)))...))--)....)))))))((((((........))))))..))).)))))) ( -45.80) >DroSec_CAF1 40846 102 + 1 ACUUUGAAGCCGCUGCCGAUGGUGAAGC------------UAUGGCCAGAGCUAAGGCUCAAGCCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGCAUCGGUCGACGGAAGUGGA .........(((((.((((((((((.((------------(.((((....)))).))))))...))))((--(((((..(((.........)))..)))))))...))).))))). ( -41.60) >DroSim_CAF1 42732 114 + 1 ACUUUGAAGCCGCUGCCGAUGGUGAAGCUAUAGCAAAAGCUAUGGCCAGAGCUAAGGCUCAAGCCCAUGA--UGAUGUGGCCAUGAACGAGGGCUGCAUCGGUCGACGGCAGAGGA .........((.((((((((((....((((((((....))))))))..((((....))))....))))((--(((((..(((.........)))..)))).)))..)))))).)). ( -50.50) >DroEre_CAF1 42310 106 + 1 ACUUUGAAGCCGCUGCCGAUGGUUAAGCUAUAGCAAAAGCUAUGGCCAGAACUAAGGCUCAAGUCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGCAUCGA--------AAGGGG .((((..(((.((((((..(((((..((((((((....))))))))...))))).)))...)))....))--)((((..(((.........)))..)))).)--------)))... ( -37.50) >DroYak_CAF1 42878 106 + 1 ACUUUGAAGCCGCUGCCGAUGGUAAAGCUAUAGCAAAAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC--UUAUGUGGCCAUGAACGAGGGCUGCAUCGC--------AGGCGG .........(((((..(((((....(((((((((....)))))(((((((((...(((....)))...))--))...))))).........)))).))))).--------.))))) ( -40.00) >DroAna_CAF1 39907 106 + 1 ACUUUGAAGCCGAUGCUGAUGCUGAUGCUGAUGCUGAUG------CUGAUGCUAAGGCCCUUGGCAAAGCCAUGAUGUGGCCAUGAACGAGGGCUGCAUCGACCGA----AAGGCA .......(((....((....((....))....))....)------))(((((...((((((((.((..(((((...)))))..))..)))))))))))))(.((..----..))). ( -37.60) >consensus ACUUUGAAGCCGCUGCCGAUGGUGAAGCUAUAGCAAAAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC__UGAUGUGGCCAUGAACGAGGGCUGCAUCGAUCGA____AGGGGA ........((....(((..((((...((((((((....))))))))....)))).)))....))........((((((((((.........))))))))))............... (-23.90 = -25.93 + 2.03)



| Location | 21,214,165 – 21,214,279 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 79.15 |
| Mean single sequence MFE | -35.40 |
| Consensus MFE | -19.58 |
| Energy contribution | -21.63 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.815555 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21214165 114 - 27905053 ACCACUUCCGUCGAACGAUCCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGGCUUGAGCCUUAGCUUUGGCCAUAGCUUUUGCUAUAGCUUCACCAUCGGCAGCGGCUUCAAAGU .((.((.(((..((((((........))))))..(((((....(--((..((((....))))..))).)))))(((((....)))))...........))).)).))......... ( -36.10) >DroSec_CAF1 40846 102 - 1 UCCACUUCCGUCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGGCUUGAGCCUUAGCUCUGGCCAUA------------GCUUCACCAUCGGCAGCGGCUUCAAAGU .......((((.(.((((((........(((.(((((((....(--((..((((....))))..))).))))))))------------)).....))))))).))))......... ( -33.62) >DroSim_CAF1 42732 114 - 1 UCCUCUGCCGUCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA--UCAUGGGCUUGAGCCUUAGCUCUGGCCAUAGCUUUUGCUAUAGCUUCACCAUCGGCAGCGGCUUCAAAGU ......(((((.(.((((((.(((....(((((.((((.(((..--..))))))))))))....)))..(((.(((((....))))).)))....))))))).)))))........ ( -40.70) >DroEre_CAF1 42310 106 - 1 CCCCUU--------UCGAUGCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGACUUGAGCCUUAGUUCUGGCCAUAGCUUUUGCUAUAGCUUAACCAUCGGCAGCGGCUUCAAAGU .((((.--------((((((........(((((((.........--.)))))))((((((..((((...(((....)))...))))..)))))).)))))).)).))......... ( -29.00) >DroYak_CAF1 42878 106 - 1 CCGCCU--------GCGAUGCAGCCCUCGUUCAUGGCCACAUAA--GCAUGGGCUUGAGCCUUAGCUUUGGCCAUAGCUUUUGCUAUAGCUUUACCAUCGGCAGCGGCUUCAAAGU ((((.(--------((((((.(((..........(((((...((--((..((((....))))..)))))))))(((((....))))).)))....)))).)))))))......... ( -38.90) >DroAna_CAF1 39907 106 - 1 UGCCUU----UCGGUCGAUGCAGCCCUCGUUCAUGGCCACAUCAUGGCUUUGCCAAGGGCCUUAGCAUCAG------CAUCAGCAUCAGCAUCAGCAUCAGCAUCGGCUUCAAAGU ...(((----(.(((((((((.(((((.(..((.(((((.....))))).)).).)))))....((....(------(....))....))..........)))))))))..)))). ( -34.10) >consensus UCCCCU____UCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA__GCAUGGGCUUGAGCCUUAGCUCUGGCCAUAGCUUUUGCUAUAGCUUCACCAUCGGCAGCGGCUUCAAAGU ..................((.((((((.(((.((((...............((((.((((....)))).))))(((((....))))).......)))).))))).)))).)).... (-19.58 = -21.63 + 2.06)



| Location | 21,214,201 – 21,214,305 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 72.74 |
| Mean single sequence MFE | -34.83 |
| Consensus MFE | -20.10 |
| Energy contribution | -20.52 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21214201 104 + 27905053 AAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGGAUCGUUCGACGGAAGUGGUGGCGUUGAGAUGGUGGGGAUUCAUGG-------- .((((.((((....)))).))))....(((((.(--((((((.(((.(((((((........)))))))((....))))).))))).))...))))).........-------- ( -36.40) >DroSec_CAF1 40874 100 + 1 ----UAUGGCCAGAGCUAAGGCUCAAGCCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGCAUCGGUCGACGGAAGUGGAGUCGGUGAGAUGGUGGGGAUUCAUGG-------- ----(((((...((((....))))...(((((.(--(((((..(((.........)))..))))))((.((....)).))(((.....))).)))))...))))).-------- ( -37.40) >DroSim_CAF1 42768 112 + 1 AAGCUAUGGCCAGAGCUAAGGCUCAAGCCCAUGA--UGAUGUGGCCAUGAACGAGGGCUGCAUCGGUCGACGGCAGAGGAGGCGGUGAGAUGGUGGGGAUUCAUGGAGGGGAUU ...((((((...((((....))))...(((((.(--(((((..(((.........)))..)))).(((..(......)..)))......)).)))))...))))))........ ( -37.40) >DroEre_CAF1 42346 96 + 1 AAGCUAUGGCCAGAACUAAGGCUCAAGUCCAUGC--UGAUGUGGCCAUGAACGAGGGCUGCAUCGA--------AAGGGGGGCGGCGGGUUGGUGGGGUUUCAUGG-------- ...((((((((...((((..((((..((((.(..--(((((..(((.........)))..))))).--------.)...))))...))))))))..)))..)))))-------- ( -32.20) >DroYak_CAF1 42914 90 + 1 AAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC--UUAUGUGGCCAUGAACGAGGGCUGCAUCGC--------AGGCGGG------GAUUGGUGGGGAUUCGUGG-------- ....((((((((((((...(((....)))...))--))...)))))))).(((((..((.(((((.--------.......------...))))).)).)))))..-------- ( -30.40) >DroAna_CAF1 39943 87 + 1 AUG------CUGAUGCUAAGGCCCUUGGCAAAGCCAUGAUGUGGCCAUGAACGAGGGCUGCAUCGACCGA----AAGGCAGACGGA-AGCUGGUGAGG---------------- ...------.((((((...((((((((.((..(((((...)))))..))..)))))))))))))).((..----..))(((.(...-.))))......---------------- ( -35.20) >consensus AAGCUAUGGCCAAAGCUAAGGCUCAAGCCCAUGC__UGAUGUGGCCAUGAACGAGGGCUGCAUCGAUCGA____AGGGGAGGCGGUGAGAUGGUGGGGAUUCAUGG________ ...((((((((...((((.(((....))).......((((((((((.........)))))))))).........................))))..))..))))))........ (-20.10 = -20.52 + 0.42)


| Location | 21,214,201 – 21,214,305 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 72.74 |
| Mean single sequence MFE | -25.40 |
| Consensus MFE | -13.29 |
| Energy contribution | -14.07 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.730432 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 21214201 104 - 27905053 --------CCAUGAAUCCCCACCAUCUCAACGCCACCACUUCCGUCGAACGAUCCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGGCUUGAGCCUUAGCUUUGGCCAUAGCUU --------.......................((.............((((((........))))))(((((((....(--((..((((....))))..))).))))))).)).. ( -27.60) >DroSec_CAF1 40874 100 - 1 --------CCAUGAAUCCCCACCAUCUCACCGACUCCACUUCCGUCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGGCUUGAGCCUUAGCUCUGGCCAUA---- --------.((((.................((((.........))))...((((..(((.........)))..)))).--.))))((((.((((....)))).))))...---- ( -27.40) >DroSim_CAF1 42768 112 - 1 AAUCCCCUCCAUGAAUCCCCACCAUCUCACCGCCUCCUCUGCCGUCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA--UCAUGGGCUUGAGCCUUAGCUCUGGCCAUAGCUU .........(((((............((((.((.......)).)).))..((((..(((.........)))..)))).--)))))((((.((((....)))).))))....... ( -26.60) >DroEre_CAF1 42346 96 - 1 --------CCAUGAAACCCCACCAACCCGCCGCCCCCCUU--------UCGAUGCAGCCCUCGUUCAUGGCCACAUCA--GCAUGGACUUGAGCCUUAGUUCUGGCCAUAGCUU --------................................--------..((((..(((.........)))..))))(--((((((.((.((((....)))).)))))).))). ( -19.40) >DroYak_CAF1 42914 90 - 1 --------CCACGAAUCCCCACCAAUC------CCCGCCU--------GCGAUGCAGCCCUCGUUCAUGGCCACAUAA--GCAUGGGCUUGAGCCUUAGCUUUGGCCAUAGCUU --------...................------.....((--------((...)))).....(((.(((((((...((--((..((((....))))..)))))))))))))).. ( -24.90) >DroAna_CAF1 39943 87 - 1 ----------------CCUCACCAGCU-UCCGUCUGCCUU----UCGGUCGAUGCAGCCCUCGUUCAUGGCCACAUCAUGGCUUUGCCAAGGGCCUUAGCAUCAG------CAU ----------------........(((-.(((........----.)))..(((((.(((((.(..((.(((((.....))))).)).).)))))....)))))))------).. ( -26.50) >consensus ________CCAUGAAUCCCCACCAUCUCACCGCCUCCCCU____UCGACCGAUGCAGCCCUCGUUCAUGGCCACAUCA__GCAUGGGCUUGAGCCUUAGCUCUGGCCAUAGCUU ..................................................((((..(((.........)))..))))........((((.((((....)))).))))....... (-13.29 = -14.07 + 0.78)
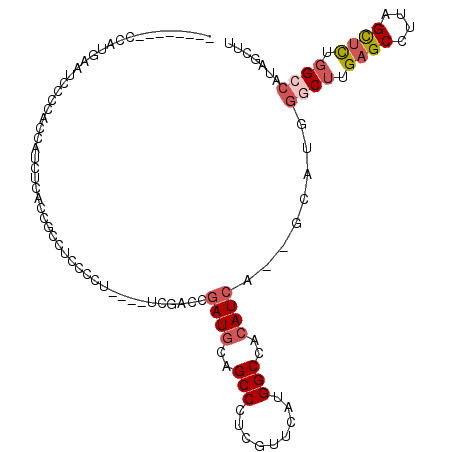



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:57:43 2006