
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 20,842,292 – 20,842,426 |
| Length | 134 |
| Max. P | 0.987646 |

| Location | 20,842,292 – 20,842,386 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 89.36 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -18.45 |
| Energy contribution | -18.32 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.22 |
| Mean z-score | -3.22 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20842292 94 - 27905053 CCUCAGAUUCAUCAAUUGGUUUCGAAUCCUCCUCGGAUUCGCCAAUUGGUUUCUUAUCCUCAGCGGAUUCAUCAAUUGGUUUCUUAUCAACCUU .................((((..((((((.....))))))((((((((((.....((((.....))))..))))))))))........)))).. ( -24.70) >DroSec_CAF1 58150 94 - 1 CCUCGGAUUCAUCAAUUGGUUUCGAAUCCUCCUCUGAUUCAUCAAUUGGUUUCUUAUCCUCAGCGGAUUCAUCAAUUGGUUUCUCAUCAACCUU ....(((((((((....)))...)))))).....(((...((((((((((.....((((.....))))..))))))))))...)))........ ( -19.40) >DroEre_CAF1 55574 94 - 1 CCUCAGAUUCAUCGAUUGGUUUCGAAUCCUCCUCGGAUUCAUCGAUUGGCUUCUUAUCCUCAGUGGAUACAUCAAUGGGUUUCCUAUCAACCUU ....((((((((((((((((...((((((.....))))))))))))))......(((((.....))))).....))))))))............ ( -21.40) >DroYak_CAF1 51706 94 - 1 CUUCAGAUUCAUCAAUUGGUUUCUUGUCCUCCUCGGAUUCAUCAAUUGGUUUCGAGUCCUCAGCGGAUUCAUCAGUUGGUUUCUUAUCAACCUU .....((...((((((((((.....((((.....))))..)))))))))).))((((((.....))))))....((((((.....))))))... ( -24.50) >consensus CCUCAGAUUCAUCAAUUGGUUUCGAAUCCUCCUCGGAUUCAUCAAUUGGUUUCUUAUCCUCAGCGGAUUCAUCAAUUGGUUUCUUAUCAACCUU .....(((..((((((((((...((((((.....)))))))))))))))).....((((.....))))..)))....((((.......)))).. (-18.45 = -18.32 + -0.13)

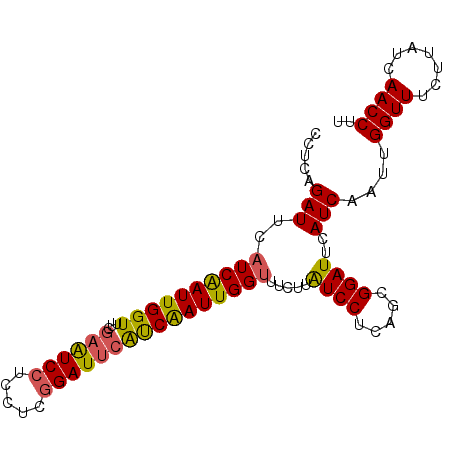

| Location | 20,842,323 – 20,842,426 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 84.14 |
| Mean single sequence MFE | -19.08 |
| Consensus MFE | -14.26 |
| Energy contribution | -14.70 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.956341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20842323 103 - 27905053 UUCUUAUCUUCUGCAGAUUCUUCAACCGGUUUCUUGUCCUCCUCAGAUUCAUCAAUUGGUUUCGAAUCCUCCUCGGAUUCGCCAAUUGGUUUCUUAUCCUCAG ..........(((..(((.........((........)).....(((...(((((((((...(((((((.....)))))))))))))))).))).)))..))) ( -22.00) >DroSec_CAF1 58181 103 - 1 UUCUUAUCUUCUUCAGAUUCUUCAACUGGUUUCUUGUCCUCCUCGGAUUCAUCAAUUGGUUUCGAAUCCUCCUCUGAUUCAUCAAUUGGUUUCUUAUCCUCAG ..............(((..((......))..)))..........((((..((((((((((...(((((.......))))))))))))))).....)))).... ( -16.70) >DroEre_CAF1 55605 103 - 1 UUCUUAUCCUCUUCAGAUUCUUCGACUGGUUUCUUGUCCUCCUCAGAUUCAUCGAUUGGUUUCGAAUCCUCCUCGGAUUCAUCGAUUGGCUUCUUAUCCUCAG ...............((.(((......((........)).....))).)).(((((((((...((((((.....))))))))))))))).............. ( -17.10) >DroYak_CAF1 51737 94 - 1 ---------UCCUCGAAAUCGUCAACUGGUUUCUUAUCCUCUUCAGAUUCAUCAAUUGGUUUCUUGUCCUCCUCGGAUUCAUCAAUUGGUUUCGAGUCCUCAG ---------..(((((((((.....((((.............)))).......(((((((.....((((.....))))..))))))))))))))))....... ( -20.52) >consensus UUCUUAUCUUCUUCAGAUUCUUCAACUGGUUUCUUGUCCUCCUCAGAUUCAUCAAUUGGUUUCGAAUCCUCCUCGGAUUCAUCAAUUGGUUUCUUAUCCUCAG ...............((.(((......((........)).....))).))((((((((((...((((((.....))))))))))))))))............. (-14.26 = -14.70 + 0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:54:31 2006