
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 20,810,356 – 20,810,447 |
| Length | 91 |
| Max. P | 0.871184 |

| Location | 20,810,356 – 20,810,447 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 80.15 |
| Mean single sequence MFE | -20.33 |
| Consensus MFE | -9.84 |
| Energy contribution | -10.34 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.871184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
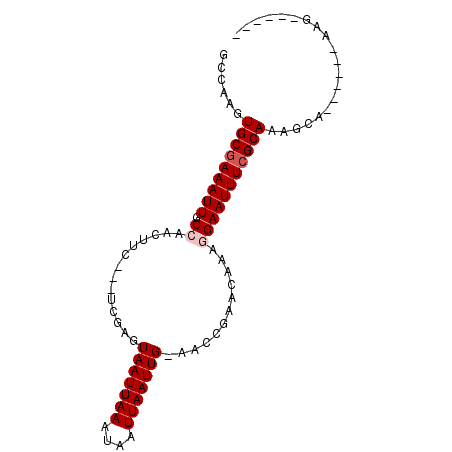
>3R_DroMel_CAF1 20810356 91 + 27905053 GCCAAGUGCGAAAUUGCCAACUUGGCGUCGAGUAAUUAAAUAAUUAAUUG-AACCGAACAAAGGAAUUUCGCAAGUCA-AAGCCAAGAAAUGC ((((((((((....)))..))))))).(((..(((((((....)))))))-...))).....((.....(....)...-...))......... ( -19.60) >DroPse_CAF1 19867 74 + 1 GCCAAGUGCGAAAUUGCCAACUUC---UGGCGUAAUUAAAUAAUUAAUUGGAAUCAAACAAGAGAAUUUCGCAAA---------AA------- ......(((((((((((((.....---)))).(((((((....)))))))..............)))))))))..---------..------- ( -17.30) >DroEre_CAF1 23619 90 + 1 GCCAAGUGCGAAAUUGCCAACUUGGCGUCGAGUAAUUAAAUAAUUAAUUG-AACUGAACAAAGGAAUUUCGCAAGGCA--AGCCAAGAAAUGC (((...((((((((((((.....))).((((.(((((....))))).)))-)............))))))))).))).--............. ( -23.80) >DroYak_CAF1 19089 92 + 1 GCCAAGUGCGAAAUUGCCAACUUGGCGUCGAGUAAUUAAAUAAUUAAUUG-AACCGAACAAAGGAAUUUCGCAAGGCAAAAGCCAAGAAAUGC (((...((((((((((((.....))).(((..(((((((....)))))))-...))).......))))))))).)))................ ( -25.00) >DroAna_CAF1 20320 77 + 1 GCCAAGUGCUAAAUUGCCAGCUUC---UGGCGUAAUUAAAUAAUUAAUUG-AACCGCACAAAGGAAUUUCGCAAAGCG------AAG------ .((..((((......(((((...)---)))).(((((((....)))))))-....))))...))...(((((...)))------)).------ ( -19.00) >DroPer_CAF1 19155 74 + 1 GCCAAGUGCGAAAUUGCCAACUUC---UGGCGUAAUUAAAUAAUUAAUUGGAAUCAAACAAGAGAAUUUCGCAAA---------AA------- ......(((((((((((((.....---)))).(((((((....)))))))..............)))))))))..---------..------- ( -17.30) >consensus GCCAAGUGCGAAAUUGCCAACUUC___UCGAGUAAUUAAAUAAUUAAUUG_AACCGAACAAAGGAAUUUCGCAAAGCA______AAG______ ......(((((((((.((..............(((((((....)))))))............))))))))))).................... ( -9.84 = -10.34 + 0.50)


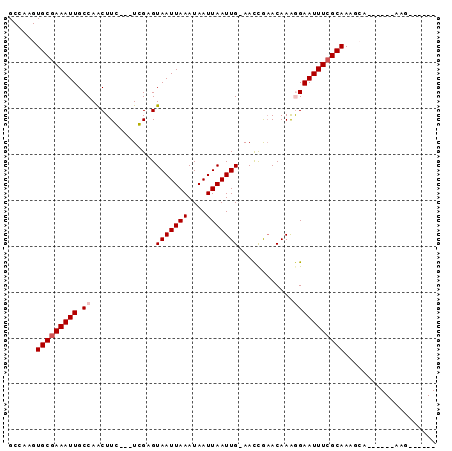
| Location | 20,810,356 – 20,810,447 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 80.15 |
| Mean single sequence MFE | -19.90 |
| Consensus MFE | -10.58 |
| Energy contribution | -10.52 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.773043 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20810356 91 - 27905053 GCAUUUCUUGGCUU-UGACUUGCGAAAUUCCUUUGUUCGGUU-CAAUUAAUUAUUUAAUUACUCGACGCCAAGUUGGCAAUUUCGCACUUGGC ..........(((.-.(...(((((((((.....(((.(((.-.((((((....))))))))).)))(((.....)))))))))))))..))) ( -20.40) >DroPse_CAF1 19867 74 - 1 -------UU---------UUUGCGAAAUUCUCUUGUUUGAUUCCAAUUAAUUAUUUAAUUACGCCA---GAAGUUGGCAAUUUCGCACUUGGC -------..---------..(((((((((...............((((((....))))))..((((---(...))))))))))))))...... ( -15.90) >DroEre_CAF1 23619 90 - 1 GCAUUUCUUGGCU--UGCCUUGCGAAAUUCCUUUGUUCAGUU-CAAUUAAUUAUUUAAUUACUCGACGCCAAGUUGGCAAUUUCGCACUUGGC ((........)).--.(((.(((((((((.....(((.(((.-.((((((....))))))))).)))(((.....))))))))))))...))) ( -23.30) >DroYak_CAF1 19089 92 - 1 GCAUUUCUUGGCUUUUGCCUUGCGAAAUUCCUUUGUUCGGUU-CAAUUAAUUAUUUAAUUACUCGACGCCAAGUUGGCAAUUUCGCACUUGGC ((.......(((....))).(((((((((.....(((.(((.-.((((((....))))))))).)))(((.....))))))))))))....)) ( -23.90) >DroAna_CAF1 20320 77 - 1 ------CUU------CGCUUUGCGAAAUUCCUUUGUGCGGUU-CAAUUAAUUAUUUAAUUACGCCA---GAAGCUGGCAAUUUAGCACUUGGC ------.((------(((...)))))...((...((((....-.((((((....))))))..((((---(...)))))......))))..)). ( -20.00) >DroPer_CAF1 19155 74 - 1 -------UU---------UUUGCGAAAUUCUCUUGUUUGAUUCCAAUUAAUUAUUUAAUUACGCCA---GAAGUUGGCAAUUUCGCACUUGGC -------..---------..(((((((((...............((((((....))))))..((((---(...))))))))))))))...... ( -15.90) >consensus ______CUU______UGCCUUGCGAAAUUCCUUUGUUCGGUU_CAAUUAAUUAUUUAAUUACGCCA___CAAGUUGGCAAUUUCGCACUUGGC ....................(((((((((((.............((((((....))))))...............)).)))))))))...... (-10.58 = -10.52 + -0.05)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:54:12 2006