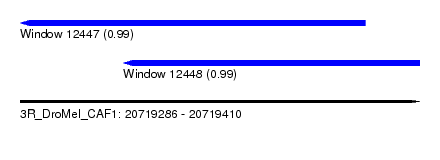

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 20,719,286 – 20,719,410 |
| Length | 124 |
| Max. P | 0.994472 |

| Location | 20,719,286 – 20,719,393 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 88.29 |
| Mean single sequence MFE | -22.14 |
| Consensus MFE | -17.66 |
| Energy contribution | -17.90 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.80 |
| SVM decision value | 2.48 |
| SVM RNA-class probability | 0.994472 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
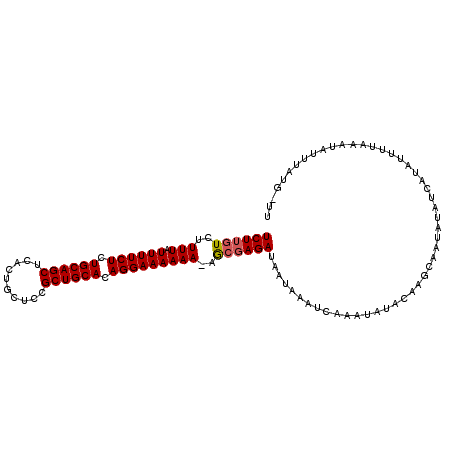
>3R_DroMel_CAF1 20719286 107 - 27905053 UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA-AGCGAGAUAAUAAAUCUAAUAUACAAGCAAUAUAUCAUAUUUUAAAUAUUUAUG-UU (((((.(((((.((((((.((((((..........)))))).)))))))))-))))))).....................((((((..((((.....)))).))))-)) ( -22.30) >DroSec_CAF1 15532 108 - 1 UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAAUAGCGAGA-AAUAAAUCUAAUAUACAAGCAAUAUAUCAUAUUUUAAAUAUUUAUCCUU ((((((..(((.((((((.((((((..........)))))).)))))))))..))))))-................................................. ( -20.70) >DroSim_CAF1 16435 108 - 1 UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAAUAGCGAGA-AAUAAAUCGAAUAUACAAGCAAUAUAUCAUAUUUUAAAUAUUUAUCCUU ((((((..(((.((((((.((((((..........)))))).)))))))))..))))))-................................................. ( -20.70) >DroEre_CAF1 16495 107 - 1 UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA-AGCCAGAUACUAAUUCAAGUAUACAAGCAAUAUUGAAUAGUUUGAAUAUUUAUG-GU (((.(.(((((.((((((.((((((..........)))))).)))))))))-))).))).....((((((((((.(((......))).))).))))))).......-.. ( -24.10) >DroYak_CAF1 16738 101 - 1 UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAACAAAGAGAUACUAAAUCAAAUAUUAAAGUAA-------UAGCUUAAAUAUUUAUG-UU ...((((((((.((((((.((((((..........)))))).)))))).....)))))))).......(((((((.((((..-------..)))).)))))))...-.. ( -22.90) >consensus UCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA_AGCGAGAUAAUAAAUCAAAUAUACAAGCAAUAUAUCAUAUUUUAAAUAUUUAUG_UU ((((((..(((.((((((.((((((..........)))))).)))))))))..)))))).................................................. (-17.66 = -17.90 + 0.24)


| Location | 20,719,318 – 20,719,410 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 90.30 |
| Mean single sequence MFE | -23.96 |
| Consensus MFE | -19.34 |
| Energy contribution | -20.62 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
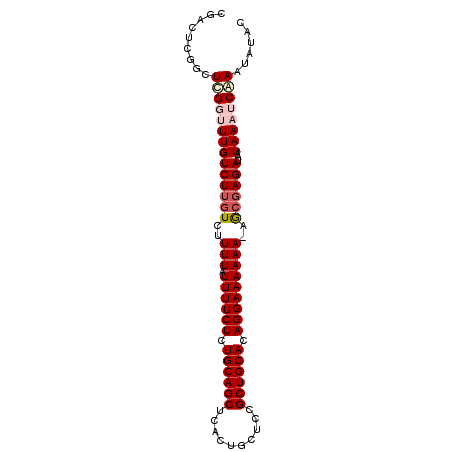
>3R_DroMel_CAF1 20719318 92 - 27905053 CCACUCGGCUCGGUUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA-AGCGAGAUAAUAAAUCUAAUAUAC ..............((((((((.(((((.((((((.((((((..........)))))).)))))))))-)))))))))).............. ( -25.30) >DroSec_CAF1 15565 92 - 1 CGACUCGGCUCGGUUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAAUAGCGAGA-AAUAAAUCUAAUAUAC ...........((((((((((((..(((.((((((.((((((..........)))))).)))))))))..))))))-..))))))........ ( -23.80) >DroSim_CAF1 16468 92 - 1 CGACUCGGCUCGGUUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAAUAGCGAGA-AAUAAAUCGAAUAUAC .........((((((((((((((..(((.((((((.((((((..........)))))).)))))))))..))))))-..))))))))...... ( -27.50) >DroEre_CAF1 16527 92 - 1 CGGCUCGGUUUGUCUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA-AGCCAGAUACUAAUUCAAGUAUAC .(((.......))).(((((.(.(((((.((((((.((((((..........)))))).)))))))))-))).)))))............... ( -22.10) >DroYak_CAF1 16763 93 - 1 CGGCUCGGUUUGUCUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAACAAAGAGAUACUAAAUCAAAUAUUA ......((((((...((((((....(((.((((((.((((((..........)))))).)))))))))....)))))).))))))........ ( -21.10) >consensus CGACUCGGCUCGGUUUGUCUUGUCUUUUAUUUUCUCUGCAGCUCACUGCUCCGCUGCACAGGAAAAAA_AGCGAGAUAAUAAAUCAAAUAUAC .........((((((((((((((..(((.((((((.((((((..........)))))).)))))))))..))))))...))))))))...... (-19.34 = -20.62 + 1.28)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:52:52 2006