
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 20,304,408 – 20,304,506 |
| Length | 98 |
| Max. P | 0.712053 |

| Location | 20,304,408 – 20,304,506 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 76.12 |
| Mean single sequence MFE | -21.75 |
| Consensus MFE | -10.23 |
| Energy contribution | -10.82 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.47 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.712053 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20304408 98 + 27905053 GCUAUAUGGUUAUAUAGCUACACAGCUAUAGUACGGCCGCACAUUCACAAAGCACCAAU-GCACACCGAGGGCAAACAAUAAACAGCAAGUGCAGAAAG .......((((.(((((((....)))))))....))))((((.........(((....)-)).........((............))..))))...... ( -19.20) >DroPse_CAF1 130503 82 + 1 GCUAUAUGGCUAUAUAUGU----AGCUACAUCGGAGCCG-------------CAGUACUCGUACACCGACGGCAAACAAUAAACAGCAAGUGCAGAAAG ((((((((......)))))----)))....((...((((-------------(.(((....)))...).))))............((....)).))... ( -17.10) >DroSec_CAF1 113241 95 + 1 GCUAUAUGGCUAUAUAGCUACACUGCUAU---ACAGCCACACAUCCACAAAGCACCACU-GCACACCGAGGGCAAACAAUAAACAGCAAGUGCAGAAAG ......(((((.((((((......)))))---).)))))..................((-((((.(....)((............))..)))))).... ( -25.10) >DroSim_CAF1 121670 95 + 1 GCUAUAUGGCUAUAUAGCUACACUGCUAU---ACUGCCGCACAUCCACAAAGCACCACU-GCACACCGAGGGCAAACAAUAAACAGCAAGUGCAGAAAG (((....(((..((((((......)))))---)..)))(........)..)))....((-((((.(....)((............))..)))))).... ( -23.70) >DroAna_CAF1 117351 95 + 1 GCUAUACAGCUAUAGAGCUAUAUAGCUAU---AUAGCUAUGGAGCUUCAAAGAGCCGCU-GCACACCGAGGGCAAACAAUAAACAGCAAGUGCAGAAAG ((((((.(((((((......))))))).)---)))))...((..((....))..)).((-((((.(....)((............))..)))))).... ( -28.30) >DroPer_CAF1 137029 82 + 1 GCUAUAUGGCUAUAUAUGU----AGCUACAUCGGAGCCG-------------CAGUACUCGUACACCGACGGCAAACAAUAAACAGCAAGUGCAGAAAG ((((((((......)))))----)))....((...((((-------------(.(((....)))...).))))............((....)).))... ( -17.10) >consensus GCUAUAUGGCUAUAUAGCUACACAGCUAU___ACAGCCGCACAUCCACAAAGCACCACU_GCACACCGAGGGCAAACAAUAAACAGCAAGUGCAGAAAG .......((((..(((((......))))).....))))......................((((.(....)((............))..))))...... (-10.23 = -10.82 + 0.59)



| Location | 20,304,408 – 20,304,506 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 76.12 |
| Mean single sequence MFE | -27.06 |
| Consensus MFE | -11.90 |
| Energy contribution | -13.90 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.647751 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20304408 98 - 27905053 CUUUCUGCACUUGCUGUUUAUUGUUUGCCCUCGGUGUGC-AUUGGUGCUUUGUGAAUGUGCGGCCGUACUAUAGCUGUGUAGCUAUAUAACCAUAUAGC .....(((((..(((((..............(((((..(-(((.(.......).))))..).))))....))))).)))))(((((((....))))))) ( -24.37) >DroPse_CAF1 130503 82 - 1 CUUUCUGCACUUGCUGUUUAUUGUUUGCCGUCGGUGUACGAGUACUG-------------CGGCUCCGAUGUAGCU----ACAUAUAUAGCCAUAUAGC ............(((((.(((((...((((.(((((......)))))-------------))))..)))))..(((----(......))))...))))) ( -19.50) >DroSec_CAF1 113241 95 - 1 CUUUCUGCACUUGCUGUUUAUUGUUUGCCCUCGGUGUGC-AGUGGUGCUUUGUGGAUGUGUGGCUGU---AUAGCAGUGUAGCUAUAUAGCCAUAUAGC ......(((((.(((((.(((((........))))).))-))))))))........(((((((((((---(((((......)))))))))))))))).. ( -38.60) >DroSim_CAF1 121670 95 - 1 CUUUCUGCACUUGCUGUUUAUUGUUUGCCCUCGGUGUGC-AGUGGUGCUUUGUGGAUGUGCGGCAGU---AUAGCAGUGUAGCUAUAUAGCCAUAUAGC ......(((((.(((((.(((((........))))).))-))))))))........((((.(((.((---(((((......))))))).))).)))).. ( -29.80) >DroAna_CAF1 117351 95 - 1 CUUUCUGCACUUGCUGUUUAUUGUUUGCCCUCGGUGUGC-AGCGGCUCUUUGAAGCUCCAUAGCUAU---AUAGCUAUAUAGCUCUAUAGCUGUAUAGC ............(((((.(((((........))))).))-)))((((......)))).....(((((---(((((((((......)))))))))))))) ( -30.60) >DroPer_CAF1 137029 82 - 1 CUUUCUGCACUUGCUGUUUAUUGUUUGCCGUCGGUGUACGAGUACUG-------------CGGCUCCGAUGUAGCU----ACAUAUAUAGCCAUAUAGC ............(((((.(((((...((((.(((((......)))))-------------))))..)))))..(((----(......))))...))))) ( -19.50) >consensus CUUUCUGCACUUGCUGUUUAUUGUUUGCCCUCGGUGUGC_AGUGGUGCUUUGUGGAUGUGCGGCUGU___AUAGCUGUGUAGCUAUAUAGCCAUAUAGC ......(((((.(((((.(((((........))))).)).)))))))).............((((((...(((((......)))))))))))....... (-11.90 = -13.90 + 2.00)


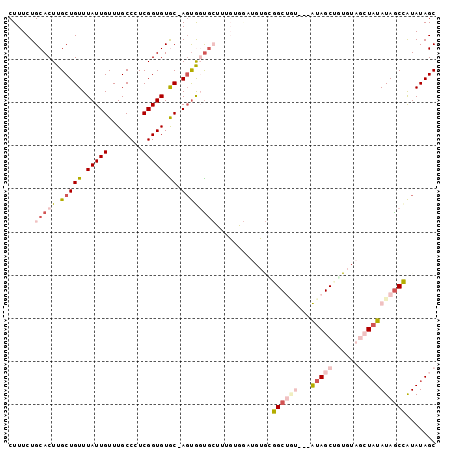
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:17 2006