
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 20,302,829 – 20,302,932 |
| Length | 103 |
| Max. P | 0.957955 |

| Location | 20,302,829 – 20,302,932 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 82.79 |
| Mean single sequence MFE | -38.26 |
| Consensus MFE | -28.88 |
| Energy contribution | -29.25 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.957955 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20302829 103 + 27905053 GGAUGGGAUGGAUGGCUGAAGGAU-----GGCCGAGGUGCUGAGGGGCGUGGCAGCUGAAGAUGGGUCAGCUAAUAAAGCGCCACUCGGUACGGUUUGCCUGGAAGCC .............((((..(((..-----((((...((((((((.(((((...((((((.......))))))......))))).))))))))))))..)))...)))) ( -41.30) >DroSec_CAF1 111654 99 + 1 ---------GGAUGGCUGAAGGAACGAGGUGCUGAGGUGCUGAGGGGCGUGGCAGCUGAAGAUGGGUCAGCUAAUAAAGCGCCACUCGGUACUGUUUGGCUGGAAGCC ---------....((((.............((..((((((((((.(((((...((((((.......))))))......))))).)))))))))..)..))....)))) ( -36.23) >DroSim_CAF1 120081 99 + 1 ---------GGAUUGCUGAAGGGCCGAGGUGCUGAGGUGCUGAGGGGCGUGGCAGCUGAAGAUGGGUCAGCUAAUAAAGCGCCACUCGGUACUGUUUGGCUGGAAGCC ---------.....(((....(((((((.......(((((((((.(((((...((((((.......))))))......))))).))))))))).)))))))...))). ( -39.10) >DroEre_CAF1 115452 87 + 1 ---------GGAUG----AAGG--------GCCGACCUGCUGAGGGGCGUGGCAGCUGAGGAUGGGUCAGCUAAUAAAGCGCCACUCGGUACAGGCUGCCUGGAAGCC ---------((...----..((--------((.(.(((((((((.(((((...((((((.......))))))......))))).)))))..))))).)))).....)) ( -36.40) >consensus _________GGAUGGCUGAAGGA______UGCCGAGGUGCUGAGGGGCGUGGCAGCUGAAGAUGGGUCAGCUAAUAAAGCGCCACUCGGUACUGUUUGCCUGGAAGCC ..............................((((((((((((((.(((((...((((((.......))))))......))))).))))))))..))))))........ (-28.88 = -29.25 + 0.38)


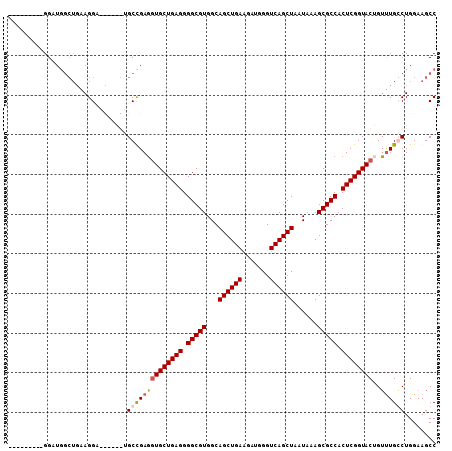
| Location | 20,302,829 – 20,302,932 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 82.79 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -19.73 |
| Energy contribution | -20.72 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916630 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 20302829 103 - 27905053 GGCUUCCAGGCAAACCGUACCGAGUGGCGCUUUAUUAGCUGACCCAUCUUCAGCUGCCACGCCCCUCAGCACCUCGGCC-----AUCCUUCAGCCAUCCAUCCCAUCC ((((...(((....(((..(.(((.((((......(((((((.......)))))))...)))).))).).....)))..-----..)))..))))............. ( -28.10) >DroSec_CAF1 111654 99 - 1 GGCUUCCAGCCAAACAGUACCGAGUGGCGCUUUAUUAGCUGACCCAUCUUCAGCUGCCACGCCCCUCAGCACCUCAGCACCUCGUUCCUUCAGCCAUCC--------- ((((...((...(((......(((.((((......(((((((.......)))))))...)))).))).((......)).....))).))..))))....--------- ( -25.30) >DroSim_CAF1 120081 99 - 1 GGCUUCCAGCCAAACAGUACCGAGUGGCGCUUUAUUAGCUGACCCAUCUUCAGCUGCCACGCCCCUCAGCACCUCAGCACCUCGGCCCUUCAGCAAUCC--------- (((.....)))....((..(((((.((((......(((((((.......)))))))...)))).....((......))..)))))..))..........--------- ( -27.10) >DroEre_CAF1 115452 87 - 1 GGCUUCCAGGCAGCCUGUACCGAGUGGCGCUUUAUUAGCUGACCCAUCCUCAGCUGCCACGCCCCUCAGCAGGUCGGC--------CCUU----CAUCC--------- ........(((.((((((...(((.((((......(((((((.......)))))))...)))).))).))))))..))--------)...----.....--------- ( -30.30) >consensus GGCUUCCAGCCAAACAGUACCGAGUGGCGCUUUAUUAGCUGACCCAUCUUCAGCUGCCACGCCCCUCAGCACCUCAGCA______UCCUUCAGCCAUCC_________ ((((.................(((.((((......(((((((.......)))))))...)))).))).((......)).............))))............. (-19.73 = -20.72 + 1.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:15 2006