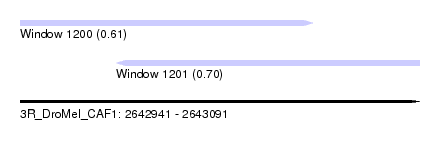

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,642,941 – 2,643,091 |
| Length | 150 |
| Max. P | 0.702242 |

| Location | 2,642,941 – 2,643,051 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.86 |
| Mean single sequence MFE | -37.10 |
| Consensus MFE | -28.51 |
| Energy contribution | -28.73 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.606654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
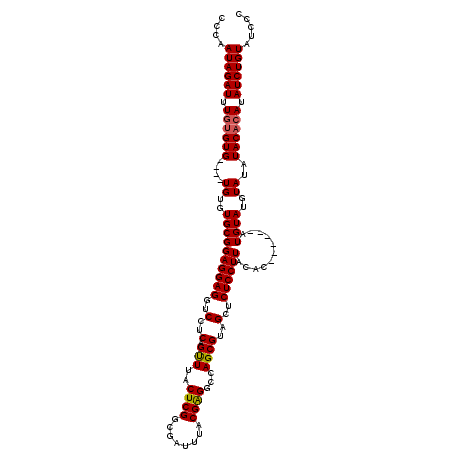
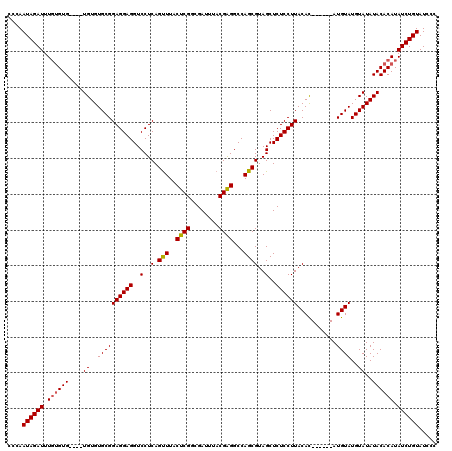
>3R_DroMel_CAF1 2642941 110 + 27905053 CCCAAUAGAUUUGUGUG----UGUGUGCGGAGGAGGUCCUCAGUUUACUCGGCGAUUUACGAGGCCAGCGUAGCUCUCCUUACAC------AUGUAUGUAUAUACACAUAUCUGUAUCCC ....((((((..(((((----((((((((((((((..(..(.(((..((((........))))...))))..)..))))))((..------..)).)))))))))))))))))))..... ( -35.20) >DroSim_CAF1 31487 114 + 1 CCCAAUAGAUUUGUGUGUGUGUGUGUGCGGAGGAGGUCCUCAGUUUACUCGGCGAUUUACGAGGCCAGCGUAGCUCUCCUUACAC------AUGUAUGUAUAUACACAUAUCUGUAUCCC ....((((((..(((((((((((..((((((((((..(..(.(((..((((........))))...))))..)..))))))....------.))))..)))))))))))))))))..... ( -37.90) >DroEre_CAF1 33996 116 + 1 CCCAAUAGAUUUUCGUG----UGUGUGCGGAGGAGGUCCUCAGCUUACUCGUCGAUUUACGGGGCCAGCGUAGCUCUCCUUACAUACGUGUAUGUAUGUAUAUACACAUAUCUGUAUCCC ....((((((....(((----((((((((((((((..(..(.(((..(((((......)))))...))))..)..))))))(((((....))))).)))))))))))..))))))..... ( -38.20) >consensus CCCAAUAGAUUUGUGUG____UGUGUGCGGAGGAGGUCCUCAGUUUACUCGGCGAUUUACGAGGCCAGCGUAGCUCUCCUUACAC______AUGUAUGUAUAUACACAUAUCUGUAUCCC ....((((((.((((((....((..((((((((((..(..(.(((..((((........))))...))))..)..))))))...........))))..))..)))))).))))))..... (-28.51 = -28.73 + 0.23)

| Location | 2,642,977 – 2,643,091 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.09 |
| Mean single sequence MFE | -37.07 |
| Consensus MFE | -31.53 |
| Energy contribution | -31.70 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.702242 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
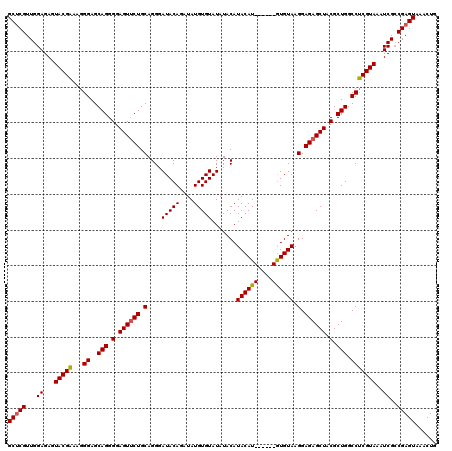
>3R_DroMel_CAF1 2642977 114 - 27905053 GCGCGUUGGAGAGUACGAAAGGGAGCAGGGGAGUCCUGCAGGGAUACAGAUAUGUGUAUAUACAUACAU------GUGUAAGGAGAGCUACGCUGGCCUCGUAAAUCGCCGAGUAAACUG ..((.((((.((.(((((......(((((.....)))))....(((((..((((((....))))))..)------))))..((..(((...)))..)))))))..)).))))))...... ( -31.50) >DroSim_CAF1 31527 114 - 1 GCUCGUUGGAGAGUACGAAAGGGAGCAGGGGAGUUCUGCAGGGAUACAGAUAUGUGUAUAUACAUACAU------GUGUAAGGAGAGCUACGCUGGCCUCGUAAAUCGCCGAGUAAACUG (((((...((...(((((...((..(((.(.((((((.(....(((((..((((((....))))))..)------))))..).)))))).).))).)))))))..))..)))))...... ( -36.00) >DroEre_CAF1 34032 120 - 1 GCUCGUUGGAGAGUACGGAAUGGAGCAGGGGAGUUCUGCAGGGAUACAGAUAUGUGUAUAUACAUACAUACACGUAUGUAAGGAGAGCUACGCUGGCCCCGUAAAUCGACGAGUAAGCUG (((((((((....(((((...((..(((.(.((((((.(....(((((....)))))...(((((((......))))))).).)))))).).))).)))))))..)))))))))...... ( -43.70) >consensus GCUCGUUGGAGAGUACGAAAGGGAGCAGGGGAGUUCUGCAGGGAUACAGAUAUGUGUAUAUACAUACAU______GUGUAAGGAGAGCUACGCUGGCCUCGUAAAUCGCCGAGUAAACUG (((((...((...(((((...((..(((.(.((((((.(....(((((....))))).......((((((....)))))).).)))))).).))).)))))))..))..)))))...... (-31.53 = -31.70 + 0.17)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:15 2006