
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,952,519 – 19,952,628 |
| Length | 109 |
| Max. P | 0.990967 |

| Location | 19,952,519 – 19,952,628 |
|---|---|
| Length | 109 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 68.66 |
| Mean single sequence MFE | -20.87 |
| Consensus MFE | -13.86 |
| Energy contribution | -13.53 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.66 |
| SVM decision value | 2.24 |
| SVM RNA-class probability | 0.990967 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
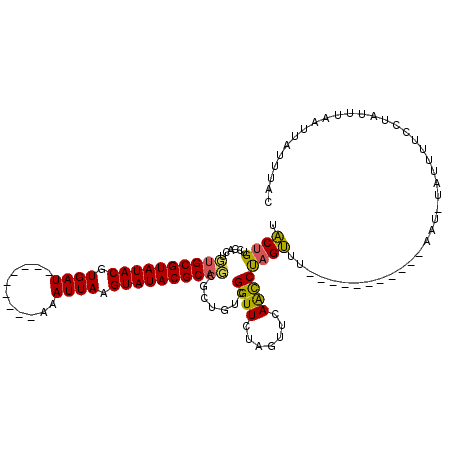
>3R_DroMel_CAF1 19952519 109 + 27905053 UACUGUCCACUGUGCGUAUACGUGAU---------AAAUUAAGUAUACGCAGCGUUGUUGGUUCUAUUUCAACCUAGUUUAUAUAUACUUGAAUUUAUUUUCAUAUUUCAUUAUUUAC ...(((..(((((((((((((.(((.---------...))).))))))))).....(((((.......))))).))))..)))......((((......))))............... ( -17.50) >DroSim_CAF1 11948 99 + 1 UACUGUCCACUCUGCGUAUACGUGAU---------AAAUUAAGUAUACGCAGCGCUAUCGGUUAUAGUUAAACCCAGUUU----------CAAUGUAUUUUCCUAUUUGAUUAUUUAC .((((....(.((((((((((.(((.---------...))).)))))))))).)....)))).((((..(((..((....----------...))..)))..))))............ ( -20.70) >DroEre_CAF1 12257 101 + 1 AGCAGUCCAUUGGGCGUAUACGUGAUUUAUUUCAAAAAUUAAGUAUACGCACUGCUGUCGGUUCUAGUUCAGUCUAGCUA-----------------UUUUCCAUUUGAAUAAUUUAC ((((((((...))((((((((.((((((.......)))))).))))))))))))))...((...(((((......)))))-----------------....))............... ( -24.40) >consensus UACUGUCCACUGUGCGUAUACGUGAU_________AAAUUAAGUAUACGCAGCGCUGUCGGUUCUAGUUCAACCUAGUUU___________AAU_UAUUUUCCUAUUUAAUUAUUUAC .((((......((((((((((.((((...........)))).)))))))))).......((((.......))))))))........................................ (-13.86 = -13.53 + -0.32)


| Location | 19,952,519 – 19,952,628 |
|---|---|
| Length | 109 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 68.66 |
| Mean single sequence MFE | -20.30 |
| Consensus MFE | -12.33 |
| Energy contribution | -13.00 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.61 |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.983010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19952519 109 - 27905053 GUAAAUAAUGAAAUAUGAAAAUAAAUUCAAGUAUAUAUAAACUAGGUUGAAAUAGAACCAACAACGCUGCGUAUACUUAAUUU---------AUCACGUAUACGCACAGUGGACAGUA ........((((.(((....)))..))))...........(((.((((.......)))).....((((((((((((.......---------.....))))))))..))))...))). ( -18.70) >DroSim_CAF1 11948 99 - 1 GUAAAUAAUCAAAUAGGAAAAUACAUUG----------AAACUGGGUUUAACUAUAACCGAUAGCGCUGCGUAUACUUAAUUU---------AUCACGUAUACGCAGAGUGGACAGUA ............................----------..((((((((.......))))....((.((((((((((.......---------.....)))))))))).))...)))). ( -23.00) >DroEre_CAF1 12257 101 - 1 GUAAAUUAUUCAAAUGGAAAA-----------------UAGCUAGACUGAACUAGAACCGACAGCAGUGCGUAUACUUAAUUUUUGAAAUAAAUCACGUAUACGCCCAAUGGACUGCU ..............(((....-----------------...((((......))))..)))..((((((((((((((...((((.......))))...))))))))((...)))))))) ( -19.20) >consensus GUAAAUAAUCAAAUAGGAAAAUA_AUU___________AAACUAGGUUGAACUAGAACCGACAGCGCUGCGUAUACUUAAUUU_________AUCACGUAUACGCACAGUGGACAGUA .........................................((((......))))..(((......((((((((((.....................))))))))))..)))...... (-12.33 = -13.00 + 0.67)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:46:55 2006